Guest blog by Dr Stephanie Suhr, Project Manager of BioMed-Bridges – a project coordinated by EMBL-EBI on behalf of ELIXIR
Say you are studying a disease in human patients, such as diabetes and obesity, and you are looking at the human DNA sequence to try and find out whether specific mutations could be contributing to it. As all mammals shared a common ancestor approximately 80 million years ago, their genomes show great similarities. This means that non-human mammals can be used as experimental models for human disease. So, to narrow down the number of suspects (disease causing mutations), you might want to compare the human genome with that of a well-established experimental model – such as the mouse – showing the same condition (phenotype): if these mice have the same mutations, this would provide an indication that the genes in question have the same function in both organisms. You have now found mouse models you can use to study the human disease, or you may look deeper into the pathways behind glucose metabolism in which these genes are involved.

Figure 1. By developing tools for data interoperability between the biological, biomedical and environmental research infrastructures, BioMedBridges provides the means to turn vast data resources into new knowledge. Image credit: EMBL-EBI, available under CC BY-ND 3.0.
However, researchers working on human patients and those working on mouse models belong to different, mostly separate communities. Each community has developed its own ways of doing things, including the terms used to describe something. For example, there are currently over 100 human genome-wide association studies (GWAS) annotated with “Diabetes” and over 750 mouse models (phenotypes) annotated with “increased circulating glucose level”. The same thing really, but how is a researcher looking through vast amounts of data to know this – and even if they do, how can they systematically compare the data?
This is what partners in the BioMedBridges project have been working on: over the last three and a half years – by developing ontologies and semantic web technologies to enable innovative, large-scale data analysis across the biological, biomedical and clinical domains – they have built the infrastructure to translate between and integrate different data sources, enabling the use of data from publicly funded research in new and different contexts. Disciplines brought together in BioMedBridges range from genomics, biological and medical imaging, structural biology, mouse disease models, clinical trials, highly contagious agents and chemical biology, to name just a few. Funded by the EC’s Seventh Framework Programme (2012 to 2016), the project’s partners consist of the communities driving 12 of Europe’s new biological, biomedical and environmental research infrastructures (Figure 1).
There is a challenge in this diversity: the different communities have different levels of understanding with respect to data integration and interoperability. To address this, BioMedBridges has taken a layered approach to data integration: interoperability is achieved via a technology stack harmonising resources across research infrastructures, which initially involves using established REST-based technology and ultimately aims to achieve more sophisticated semantic interoperability. The final step in the construction of this information infrastructure is the implementation of Web Services-based simple object queries. Once completed, this will make it possible for researchers to find – in one simple step – information most relevant to a scientific question or related to a specific disease across a very large number of data resources with millions of data entries (Figure 2).
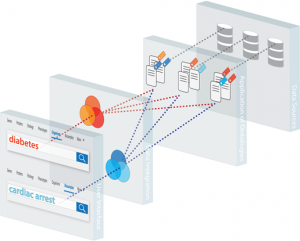
Figure 2. The information infrastructure created by BioMedBridges will contribute to enabling users to search for all items related to a specific disease across a potentially very large number of resources. Image credit: Julie McMurry/EMBL-EBI, available under CC BY 3.0.
Using this stepwise, layered approach ensures that all research infrastructures and data resources can systematically be brought to a higher level of integration. Almost as a side effect, this creates the necessary expertise of all involved to further advance data interoperability in future efforts. Ultimately, the BioMedBridges project is building a shared data culture in the life sciences, finding technical solutions to the flow of information from basic research into medical and environmental applications.
Among a host of tools that demonstrate the potential for gaining new knowledge from integrating such diverse data sources, one of the most important project outcomes may be the lessons learned. None of these are revolutionary, and some may be very obvious – for example, any effort in integrating different data sources must be driven by specific use cases: it is pointless to develop infrastructure that is not useful for research – but there is a friction in that such use cases tend to be very specific where, ultimately, generic tools may be more useful for a wider community. If anything, the size and ambition of the project provides new emphasis for both the dos and don’ts of data integration at a large scale.
The biomedical sciences research infrastructures behind BioMedBridges remain in a unique position to drive these efforts further. When BioMedBridges ends in December 2015, its follow-on project – CORBEL – will already be underway. The involvement of the research infrastructures does not only contribute to the sustainability of the tools, resources and services developed within the project, their continued collaboration also provides the opportunity to achieve real integration – possibly even a change of culture – across the life sciences with respect to data.
The symposium Open bridges for life-science data – to be held on 17-18 November 2015 on the Wellcome Trust Genome Campus in Hinxton near Cambridge, UK – will provide an opportunity for the big – and growing – community to discuss real-life challenges connected to data sharing and interoperability in the life sciences. Topics covered include semantic technologies, translational research infrastructure, APIs and workflows, ethics and security requirements for sensitive data, and metagenomics. Conference workshops include a session for researchers on journal data policies, in which the Scientific Data team is participating. The symposium provides the perfect opportunity to network, discuss and collaborate with other like-minded individuals, including leaders in the field.
Registration is open now – we hope to see you there!
Disclosure: Nature Publishing Group is represented on the Industry Advisory Committee of ELIXIR.