A cluster of pancreatic cancer cells. Scanning electron micrograph{credit} Anne Weston, LRI, CRUK. https://wellcomeimages.org{/credit}
Pancreatic cancer is a highly heterogeneous disease that often has a poor prognosis. Development of drugs or treatment strategies to target cancers, including pancreatic cancer, depends on identifying the drivers of disease. These are the genes that promote carcinogenesis and coordinate development of the cancer. But by the time a patient is diagnosed, it can often be very difficult to tell which of the many mutations present in the tumor are actually disease drivers, and which are just along for the ride.
A new paper published in Nature Genetics describes a strategy for finding the genetic drivers in pancreatic cancer. The authors used a forward genetic screen in mice that targets a particular transposable element, the piggyBac transposon, to the pancreas. When the transposon inserts itself into the genome, it disrupts genes, causing mutations that may then lead to cancer. By using the screen in “sensitized” mice (i.e., mice with particular mutations that will accelerate disease progression), the authors were able to cause pancreatic tumors to form in the mice. The genetic changes in these tumors were then examined to identify which genes are most often targeted by the transposon.
Other studies have been published recently that use a similar approach to find drivers of other types of cancer. Neal Copeland, Nancy Jenkins and colleagues pioneered the use of Sleeping Beauty transposon mutagenesis to screen for genes important in cancer, including a recent study in liver cancer associated with hepatitis B. Rama Khokha and colleagues recently used the Sleeping Beauty mutagenesis method to identify driver genes responsible for the formation of sarcomas.
These screens have been very successful; there have even been Sleeping Beauty screens for pancreatic cancer driver genes (here and here). However, Roland Rad and colleagues found that a Sleeping Beauty transposon screen was not ideal for studying certain types of pancreatic cancer. In addition, Sleeping Beauty and piggyBac have different insertion preferences, so the tools complement one another. This means that, while some sets of genes identified with the two methods do overlap, there are other genes that can only be found by using one or the other method. Importantly, Dr. Rad and colleagues observed different histological subtypes of pancreatic cancer in mice when using piggyBac, which were not observed using Sleeping Beauty.
We asked Dr. Rad, one of the lead authors of the study, to tell us a little more about the paper.
For readers unfamiliar with insertional mutagenesis screens, could you tell us what a piggyBac transposon is and how it was discovered?
Transposons are mobile DNA segments that can move around the genome. They were first discovered by Barbara McClintock more than 50 years ago. The DNA transposon piggyBac encodes a transposase, which moves the transposon from one genomic locus to another by a cut-and-paste mechanism. Transposable elements, which have been widely used for genetic screening in bacteria, yeast, arthropodes and nematodes, had been inactivated during vertebrate evolution and were hence not available as genetic tools in higher organisms until recently. Successful efforts over the past ten years to make piggyBac work in mammalian cells motivated us to target it to the mouse genome and test its applicability for somatic mutagenesis in mice.
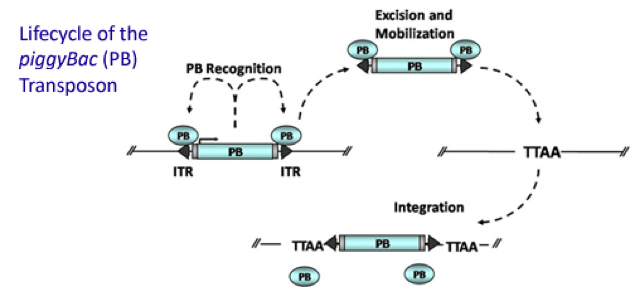
The PB transposase recognizes the specific inverted terminal repeats (ITRs) at each end of the transposon. PB then “cuts” the transposon out of its original location and moves it to a new, random location in the genome with a TTAA sequence. {credit}Transposagenbio via Wikimedia Commons{/credit}
How do screens like this (performed in mice) inform us about human cancer? What is the advantage of this approach over direct sequencing of patient tumors?
Genetic screening and cancer genome sequencing are highly complementary approaches. Sequencing and array-based analyses of patient tumors can very accurately identify all classes of somatic alterations in cancer. However, many of these changes are difficult to interpret. For example, hundreds or even thousands of genes are found to be transcriptionally or epigenetically dysregulated within a single patient´s tumor, meaning that they are not mutated but just being turned on or off. Pinpointing the few cancer-causing events among these large gene sets is extremely difficult. Likewise, copy number variation in cancer often affects large chromosomal segments, and for 75% of commonly amplified or deleted regions in human cancer, the cancer-causing genes have not yet been identified.
PiggyBac screening can tremendously facilitate this “search for the needle in the haystack” because transposons jump directly into the relevant genes. Even if a cancer gene is unequivocally identified through sequencing (for example based on its mutation), understanding downstream complexity can be difficult. Many cancer genes (e.g. methyltransferases, histone modifying enzymes, DNA repair genes) have large numbers of targets. Others (e.g. Ras) have many effector pathways that are used differently in various cancer types or have numerous interaction partners. Here again, unbiased genetic screening can identify ‘players’ at all levels of these cascades and can directly pinpoint important downstream effectors. Moreover, genetic screening provides a first level of biological validation of cancer genes and functional insights at an organismal level. These are some examples, which show that transposon-based screening can answer biological questions that cannot be systematically addressed by other approaches to cancer genome analysis.
What was the most surprising aspect of this study?
The screen produced numerous unexpected results. This is the beauty of a hypothesis-free forward genetic approach. We have discovered a large set of novel transcription factors involved in pancreatic cancer and shown that transposons can be used to identify cancer-relevant non-coding regulatory regions in the genome. The study also showed that insertional mutagenesis can induce different subtypes of pancreatic cancer and can dissect underlying genetic causes.
What was the biggest challenge your group faced during the course of the study?
The biggest challenge was to make the system work in mice. PiggyBac originates from Trichoplusia ni, the Cabbage moth. We modified PiggyBac and introduced it into the mouse genome. Naturally, we did not have a priori knowledge as to how the system would behave in the mouse. Will it be efficient enough to achieve transposition? How many transposons per cell will we need to achieve tumor induction in individual tissues? Do high transposon copies induce toxicity? How will the genetic elements (enhancers, gene trapping elements etc.) affect the phenotype? We addressed these questions by developing many different transposon mouse lines and systematically exploring PiggyBac’s characteristics in vivo.
How do you see your results being used in the future by other researchers or clinicians working with pancreatic cancer?
The study has produced rich biological insights and large sets of putative novel “players” in pancreatic cancer. Researchers will use this knowledge and take individual aspects further, e.g. perform in depth analysis of individual genes discovered in our screen or test whether they are targetable. Our genome-wide screen adds further pieces to pancreatic cancer´s “puzzle” in order to better understand the complexity of the biological processes driving tumorigenesis. We hope that this will ultimately help guide the development of novel therapeutic strategies.
You can find the paper describing this study here. More information about Dr. Rad and the piggyBac transposon system can be found here.
