A year ago now, we launched a trial to test the use of cloud-container platforms for peer review and publication of code at several Nature journals. The trial phase of this initiative is now officially over, and we would like to share the experience and outcomes, and provide an overview of what comes next.
What problem are we trying to solve?
Our guiding principle is that when new code is central to the main claims made in the paper, it is imperative that the code meets the same quality and reproducibility standards as the paper itself. This means the code needs to be properly documented, evaluated by experts so that it is functional (ie peer reviewed), and permanently identified and accessible at the time of publication to ensure the reproducibility of the results (these same principles apply to other research objects like data and protocols, but those were not the focus in this particular trial).
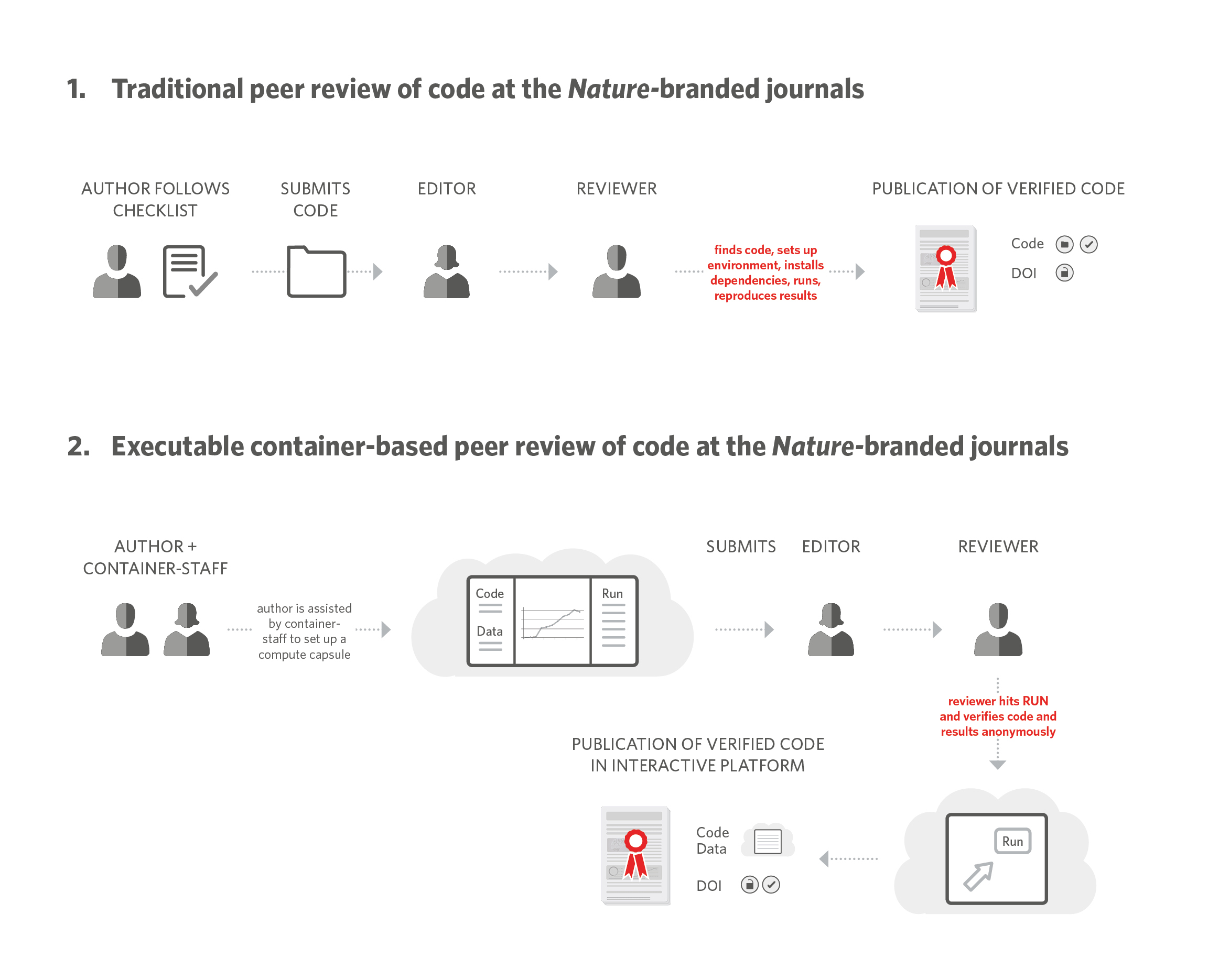
Nature Methods adopted the practice of ‘peer reviewing code’ for software papers in 2007 (editorial). Under this practice, editors require authors to submit the source code, a test dataset and details of installation, and ask the reviewers to install and test-run the code during peer review. This form of peer review is highly time consuming for authors, editors and reviewers, but it is also necessary. It is not uncommon for reviewers to point to basic flaws in the instructions or files that could render the code completely unusable.
Over a year ago, we partnered with Code Ocean, a Docker-based platform that allows authors to deposit code and data and enables users to run the code on the cloud with the set parameters to reproduce the results, or execute the code with new input values. Together we developed a set of workflows and basic functionality of the platform that enables authors to upload the code and data associated with their submission, and reviewers to access the platform anonymously during peer review (see figure, reproduced from reference 1).
The trial was meant to evaluate if such a platform would provide:
- A service to authors by assisting them in depositing the code and data and compiling them in an open, executable-based platform.
- A service to reviewers by making code peer review easier (as easy as clicking a button). Reviewers can evaluate the code in the cloud using computing time that we provide as a publisher, not their own.
- A service to readers by providing the code associated with the paper in a way that is properly identified, documented and supplied in a publically accessible platform that allows running, reusing and repurposing the code.
The trial was optional for authors at the three participating journals (Nature Methods, Nature Biotechnology and Nature Machine Intelligence) and we tracked feedback from authors and reviewers, author opt in rates and user-engagement metrics.
What have we learnt? Results!
Over 95 papers have now participated in the trial and more than 20 published papers are providing open, verified, properly documented and cited code using the technology.
Despite the additional work that authors need to do upfront when they sign up to the trial, we’ve seen large author uptake, with 54% of authors across all journals opting in to participate. Importantly, our reviewers actively engage with the platform. Capsules have received an average of 34 views via the private links provided to the reviewers. Approximately half of the reviewers signed up and duplicated the capsules, a requirement for running the code. Each reviewer that signs up runs the code 1.3 times, on average. Importantly, peer review of code in this manner has surfaced problems with some manuscripts that would have led to the ‘irreproducibility’ of the code and the results.
Upon publication, we provide the links to access the code and data in a ‘Code Availability’ statement of the paper, which is provided openly to all readers regardless of access status.
We are looking at ways to improve the workflows and experience by providing better information, an easier workflow for editors and authors and better ways of surfacing code that is shared openly and peer reviewed through the use of badges.
We have been very pleased to see that the high standards we are applying to ensure open science and reproducibility of code in our papers has been noted, as we’ve received very positive feedback about the initiative from authors, reviewers and the science community. You can read more about the initiative and the results in the below editorials and in the Science Editor piece that we recently published.
Science Editor: Three approaches to support reproducible research
Nature Biotechnology: ‘Changing coding culture’
Nature Machine Intelligence: ‘Sharing high expectations’
Nature Methods: ‘Easing the burden of code review’
What’s next?
Given the positive effects we’ve seen so far, we will continue the current practice at the journals. We also want to learn how the workflow would scale and to test it on more scientific disciplines so we have added Nature, Nature Protocols and BMC Bioinformatics to the trial.
A huge thanks to our authors, editors and reviewers who have engaged with us in this journey, we couldn’t have done it without you! We hope that this initiative, alongside others that promote data and protocol sharing, will help us develop our articles to live by the promise of more open and reproducible science.
References
1. Pastrana, E., Kousta, S. & Swaminathan, S. Three Approaches to Support Reproducible Research. Science Editor
AUTHORS: This guest blog comes from Erika Pastrana, Editorial Director for the Nature Research Journals and Sowmya Swaminathan, Head of Editorial Policy and Research Integrity at Nature Research.