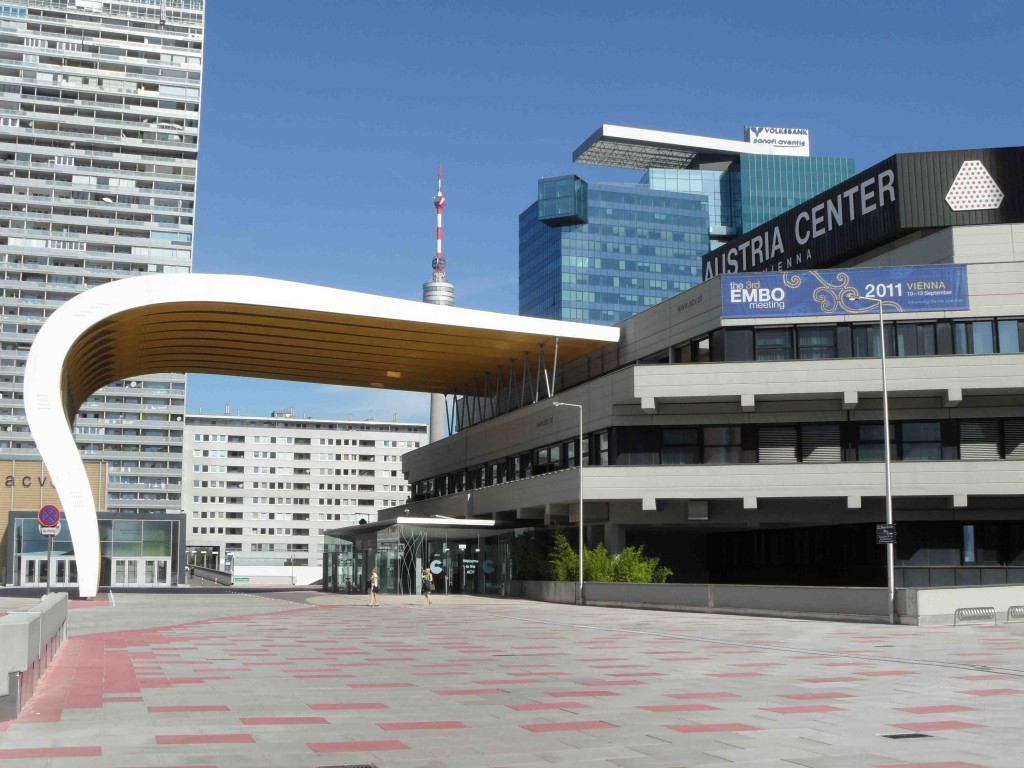
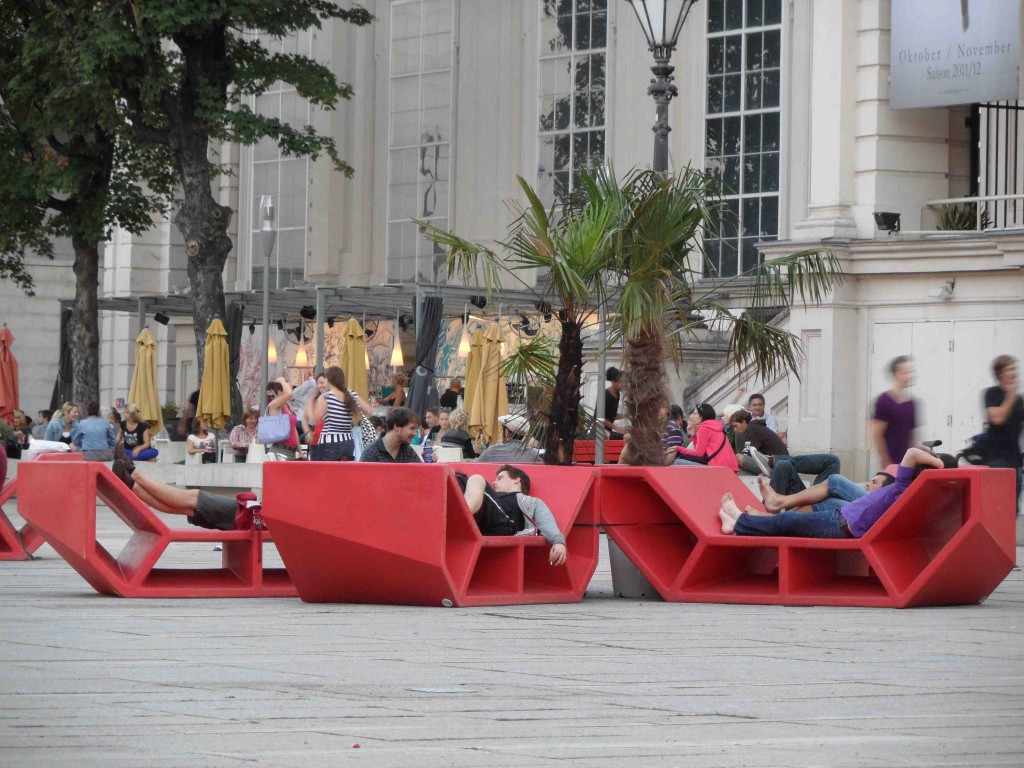
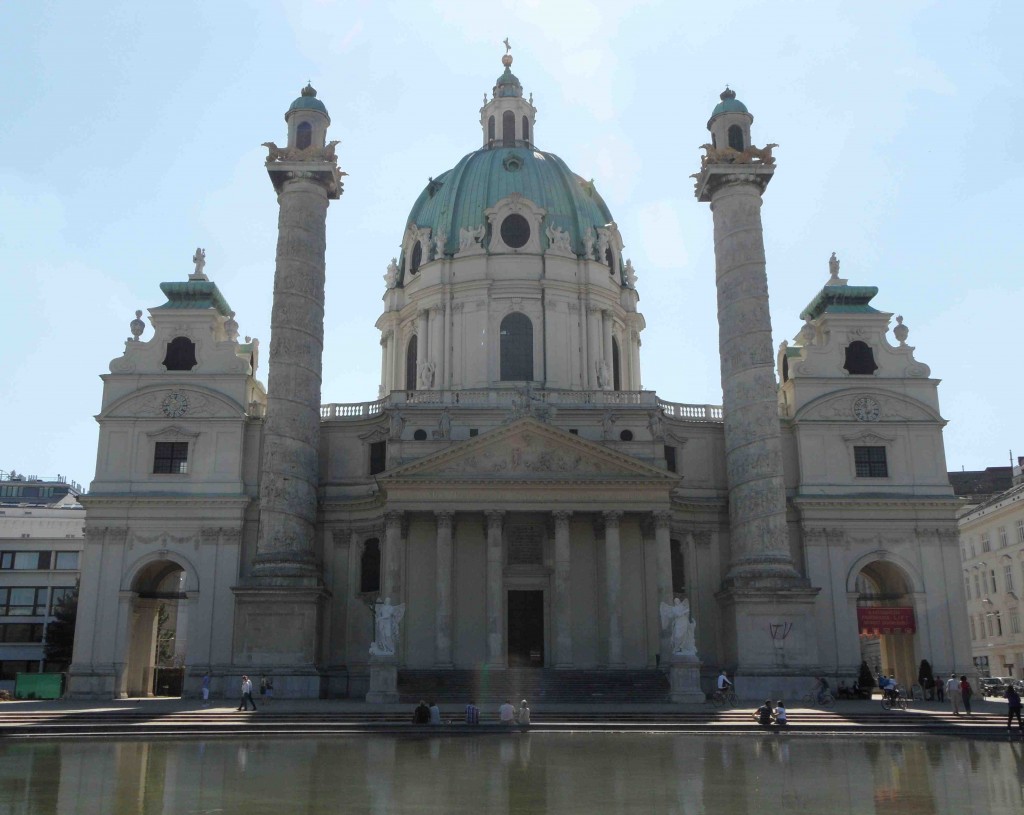
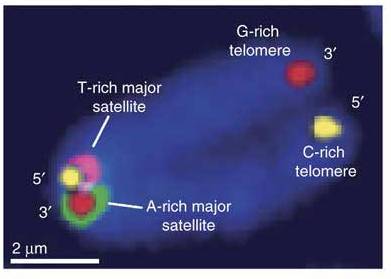
At the end of March, Dot and I spent two days in Edinburgh at the Select Biosciences meetings held at the Edinburgh Conference Centre on the Herriot-Watt campus, about 6 miles outside the city centre. The weather was record-breakingly hot for March (and Scotland!) – here is some evidence of the spring sunshine:
We didn’t have too much time to enjoy the weather though as there were four concurrent meetings to attend; I spent almost all of my time at the Single Cell Analysis Europe conference, while you will receive further reports from Dot on talks from the Lab-on-a-chip European Congress, Advances in Microarray Technology, and Advances in Biodetection & Biosensors sessions.
As an editor, I found the meeting really interesting, with some of the top names in the field speaking and methodological advances a-plenty for us to learn about! The exhibition hall was also a really interesting addition to the main sessions as we got to see a lot of the lab-on-a-chip and microfluidic equipment that people use in their protocols in real life for the first time! And they really are quite amazing!
Day one’s session on using microarrays and chips in single cell analysis started with a keynote presentation from Professor Marcus Textor from ETH Zurich, on the use of microfabricated cell culture platforms to study cell function and drug response in engineered 2D and 3D microenvironments. Describing how cell culture has moved on from the invention of the Petri dish in 1877, Professor Textor firstly showed how PDMS microwells can be engineered to provide 3D culture environments with varying substrate stiffness and shape, which can regulate the assembly of the actin cytoskeleton, impacting cell survival, phenotype and function. He also described the use of a PEG-based microwell platform to create 3D breast cancer models to allow responses to taxol and other anticancer drugs to be tested in vitro.
Dr. John Collins (NanoInk Inc.) then presented a talk on dip pen nanolithography as a technique for fabricating cellular microenvironments that allows single cell co-culture, and targeted delivery of agents such as drugs. As a work in progress, Dr. Collins also explained how they are using this technology to print substrates in lines rather than dots to enable the study of cell motility in this 3D microenvironment.
The next session, on label-free routes to single cell analysis started with an interesting talk from Professor Susann Müller (Helmholtz Centre of Environmental Research), which focused on assessing microbial community dynamics using flow cytometry and phylogenetics, encompassing analysis of even unculturable bacterial strains. Changes in the community cytometric fingerprint or structure over time can be measured and used as a biosensor for the state of natural environments, such as monitoring the stability of the enhanced biological phosphorous removal (EBPR) process in wastewater treatment plants as illustrated in this talk.
The afternoon began with the second keynote presentation, from Professor Nicholas Navin of the MD Anderson Cancer Center, which detailed the investigation of genome evolution in breast cancer by single cell sequencing. Combining flow-cytometric sorting, whole genome amplification and next-generation sequencing, this single nucleus sequencing (SNS) method can be used to accurately quantify copy number in single nuclei. Professor Navin’s group used SNS to analyse 100 single cells from a polygenomic tumour and 100 single cells from a monogenomic primary tumor and its liver metastasis, data from which indicates that tumors grow and evolve by punctuated clonal expansions, rather than gradual tumor progression. Look out for the upcoming Nature protocol on this method as we have this in production as we speak!
New and improved methods for DNA sequencing are clearly an up-and-coming area of methodological research and before moving on to the use of microfluidics in single cell analysis, the afternoon’s session continued with a talk from Professor David Bensimon (Ecole Normale Supérieure) describing a novel, single molecule method for the mechanical sequencing of DNA. This method uses a magnetic trap to mechanically open out and close DNA hairpins. In the presence of hybridizing complementary oligonucleotides, rehybridization of DNA hairpins is blocked, and the position of these roadblocks on single molecules can be measured with nanometer precision, allowing sequencing of the DNA molecule by hybridization or ligation:

For further details on this new method, see the paper just published in this issue of Nature Methods (9(4), p367-372) [Update: this paper is now free to access for a week] and the News & Views piece by Sten Linnarsson in the same issue, from which the above illustration is taken. Another method for sequencing was also presented at the same time in the lab-on-a-chip meeting, using nanopore sensors for next generation DNA sequencing. Sadly my conference-going skills do not extend to splitting myself in half to be at two talks at once, so I was unable to hear about this new method in person as well!
Day two started with two talks on the use of RT-qPCR in single cell analysis, from Professor Mikael Kubista (TATAA Biocenter) and Dr. Ken Livak (Fluidigm). These highlighted the variation in gene expression levels found from cell-to-cell, even in seemingly homogeneous populations, caused by the stochastic nature of transcription that occurs in the cell in bursts. Both speakers presented methods that can be used to deal with this heterogeneity, the latter explaining why single cell data needs to be treated differently from conventional qPCR data, and how this can be achieved using multivariate analysis and considering factors such as replicates, limits of detection, normalization and data display.
A couple more very interesting methodological talks followed in the afternoon session on single cell analysis in signalling, from Professor Ola Soderberg (Uppsala University) and Frederik Fritzsch (Dortmund University). Professsor Soderberg described the application of the in situ proximity ligation assay (PLA) and padlock probes to visualize signal pathway activity in single cells. In situ PLA utilizes pairs of antibodies, with bound DNA sequences, to target interacting proteins. Proximal binding of the antibodies and the conjugated oligonucleotide sequences creates a circular DNA molecule that can be amplified by rolling circle amplification (RCA) and subsequently detected by hybridization of fluorophore-labelled probes. Padlock probes also use RCA to generate signals for detection and the two can be combined to allow multiplexed analyses. The Envirostat 2.0 was then presented by Frederik Fritzsch with plenty of impressive videos to demonstrate this new, negative dielectrophoresis-based system for contactless single cell isolation, cultivation and analysis.
All in all, the full program provided many stimulating talks in areas undergoing, and with great potential for, methods development. Although we cover protocols in these areas already, we are now full of new ideas for enhancing our content on these topics going forward, so watch this space!