Life science research increasingly runs on software. A good fraction, perhaps even most of it, is made by academics, for academics: Rough around the edges, perhaps, but effective — not to mention free. But, is it of high quality?
Alexandros Stamatakis decided to find out.
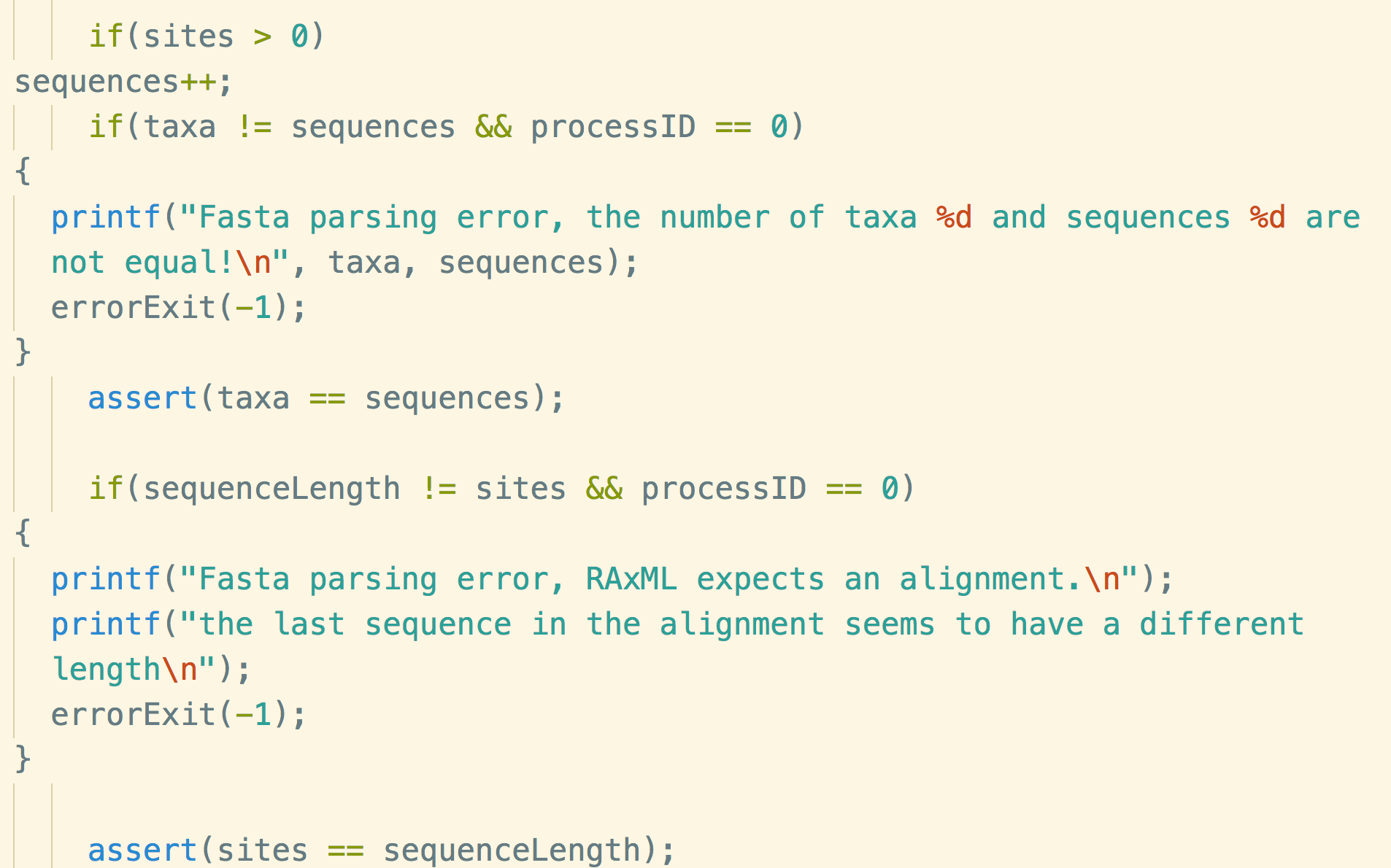
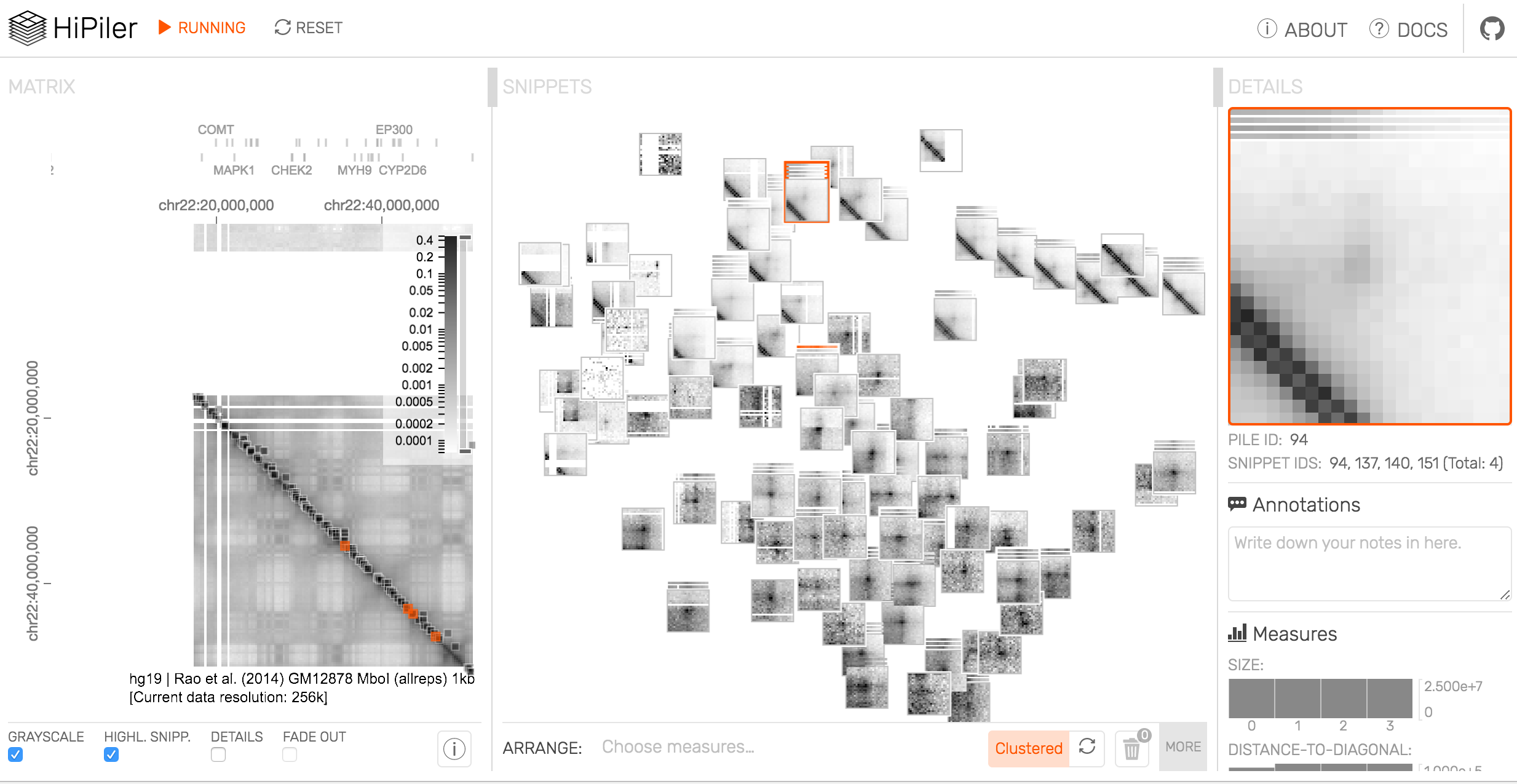
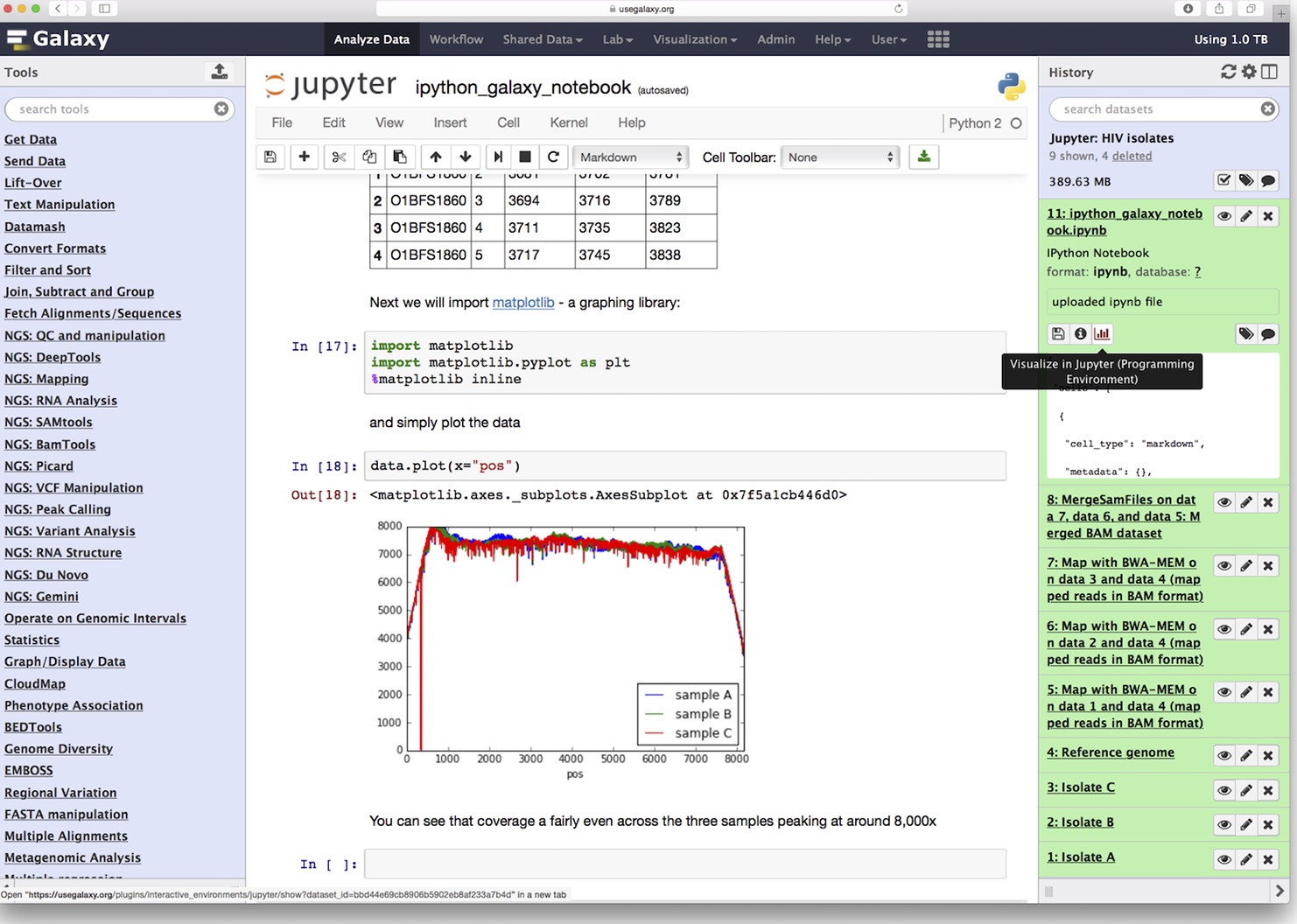
Stamatakis is a computer scientist and bioinformatician at HITS, the Heidelberg Institute for Theoretical Studies in Germany, and a professor of computer science at the Karslruhe Institute of Technology. His team has been developing and refining software tools for evolutionary biology for more than 15 years, he says, including one called RAxML (from which the code snippet shown above was pulled). Yet for all that time, he says, his code still wasn’t perfect.
“The more I developed it the more bugs I had to fix and the more I started worrying about software quality,” he says.
Not software ‘accuracy’, mind you — when it comes to phylogenetics, it’s difficult to know whether software is providing the correct answer. “You don’t know the ground-truth,” Stamatakis says. Rather, he was curious whether popular tools meet computer-science standards for quality.
To find out, Stamatakis and his team downloaded the code for 16 popular phylogenetic tools (plus, as a control, one from the field of astronomy), which collectively have been cited more than 90,000 times. They then ran those codes — 15 of which were written in C/C++ and the last in Java — through a series of tests.
For instance, they looked at how well software can scale from a desktop computer to a large cluster, something that increasingly is necessary as life science datasets balloon in size. They measured the amount of duplicated code in the software to get a rough indication of maintainability. And they counted the number of so-called ‘assertions’ — logical statements in the code that assert, for instance, that a value falls within a certain range, and that cause the software to terminate should they fail — to obtain a measure of code ‘correctness’.
“There have been empirical studies by computer scientists working in the field of software engineering, where they showed that there is a correlation between incorrect code, or code defects, and the number of assertions used — or let’s better say, an anti-correlation,” Stamatakis says.
So, how did the toolset do? Not too well.
As documented in an article published 29 January in Molecular Biology and Evolution, none of the 16 programs in the round-up, including Stamatakis’ own RAxML, aced all the tests. (With 57,233 lines of code, RAxML exhibited both compiler warnings and memory leaks.) But, he stresses, that is neither to denigrate the programmers who wrote those tools — who, after all, were simply trying (and generally succeeding) to solve a particular problem — nor to suggest they do not work properly.
Rather, he says, potential users must exercise caution in using these tools. “They shouldn’t blithely trust software. And they shouldn’t view it as black boxes,” but instead (as he puts it in his article) as “potential Pandora’s boxes”.
Users should strive also to understand what their code is doing, Stamatakis advises. And if unexpected results arise, repeat them using a separate tool that performs the same task, to ensure they aren’t chasing digital phantoms.
Stamatakis concludes his article with a series of ‘best practices’ for software developers. These include running tests for memory allocation errors and leaks, using assertions, checking for code compilation warnings using multiple compilers, and minimizing code complexity and duplication — practices that are common in professional software development but less so in the life sciences.
The tools Stamatakis’ team used to run its tests are freely available, so readers can try them themselves to see how trustworthy their chosen software is.
Journal editors, he says, should consider requiring such tests of any peer-reviewed work, either performed by the authors themselves prior to submission, or by the peer-reviewers. In fact, during our conversation, Stamatakis suggested he might make the toolbox available as a Python script or Docker container, to make it easier for others to adopt. If and when he does, we’ll let you know. In the meantime, caveat emptor!
Jeffrey Perkel is Technology Editor, Nature
Suggested posts
‘Manubot’ powers a crowdsourced ‘deep-learning’ review
eLife replaces commenting system with Hypothesis annotations