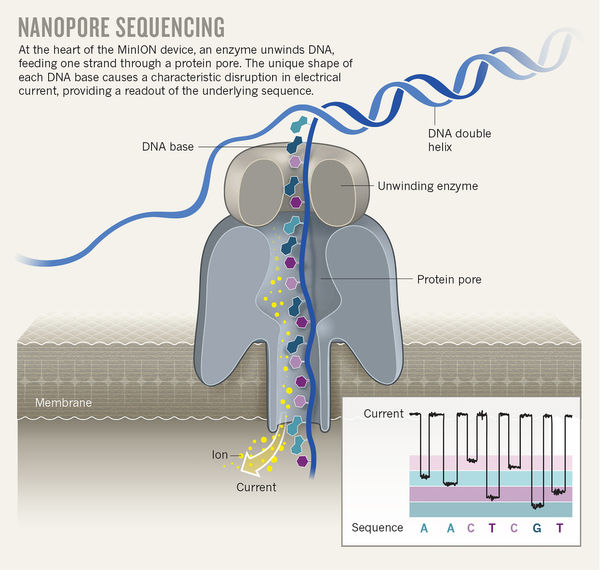
For this week’s Technology Feature, Michael Eisenstein wrote about the technology, applications, and challenges of nanopore DNA sequencing. In brief, the technology involves threading intact pieces of DNA through a tiny aperture in a membrane or other barrier, through which a current flows. As each base passes, it disrupts that current in a characteristic way, allowing specialized software to determine the sequence.
The technology has multiple benefits: it’s relatively inexpensive and compact, and produces exceptionally long reads. But the resulting error rate is also higher than some other technologies. What that means is, informatics tools designed to handle short-read data can often stumble when confronted with nanopore sequences. But a growing collection of dedicated long-read tools is rapidly filling in the gap. I asked a few nanopore veterans to help me compile a list.