Fly illustrations from The University of Texas Publication No. 4313: April 1, 1943 and The University of Texas Publication No. 4445: December 1, 1944
A paper published online this week in Nature Genetics mapped the enhancer regions of 5 fruit fly species to better understand the evolution of regulatory DNA.
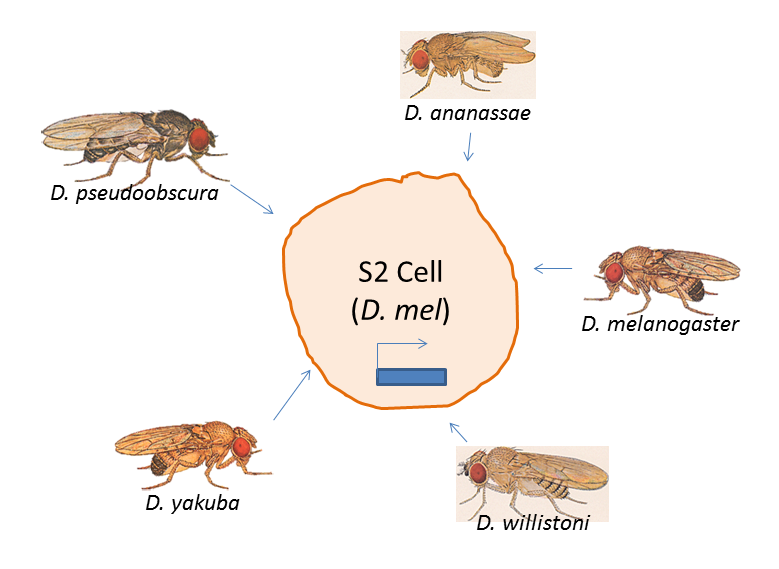
Alexander Stark and colleagues used a recently-developed method, called STARR-Seq, to find which Drosophila melanogaster enhancer elements were still functional in the different fly species. Basically, you chop up your input DNA, put the fragments into a vector with an open reading frame preceding it (so your input DNA can act as an enhancer, if it so chooses) and then toss it into some cultured cells.
In this case, the cells used were Drosophila melanogaster S2 cells. Keeping the cell line constant ensured that any differences seen in the expression levels of the ORFs + enhancers would be due to cis changes and not trans ones (like different transcription factors).
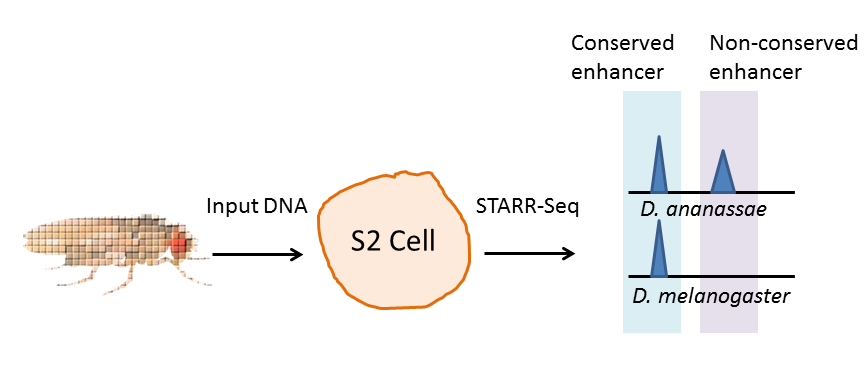
After expressing the constructs in S2 cells, you sequence the transcripts and compare them to the input and to the genomic sequence of the reference species, D. melanogaster. Interestingly, the authors found a pretty high proportion of enhancer elements are conserved between species. Between D. melanogaster and it’s closest relative used in the study, D. yakuba (only 11 million years diverged), 58% of the D. melanogaster elements were conserved. Between the most distant relatives (D. mel and D. willistoni), 34% were conserved. Now, they may just look like flies to you and me, but those two species are about as distantly related as you and I are from lizards.
Another key finding was that even over relatively short evolutionary time, hundreds of new enhancers can appear, right out of the blue. DNA sequences that had previously done nothing (or at least, done something completely different) were transformed into working enhancers. Between D. mel and D. yak, D. mel gained 525 enhancers, while its yellower relative gained 472.
As for losses, the authors estimated that every 10 million years, about 4% of enhancers lose their activity. This rate of gain and loss of enhancer elements is probably faster than was previously thought. The authors speculate that the rates are likely to be much higher in mammals. Another example of why regulatory DNA is so important to the evolution of gene expression and function.