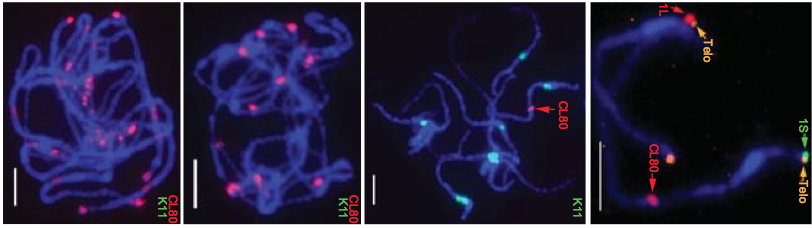
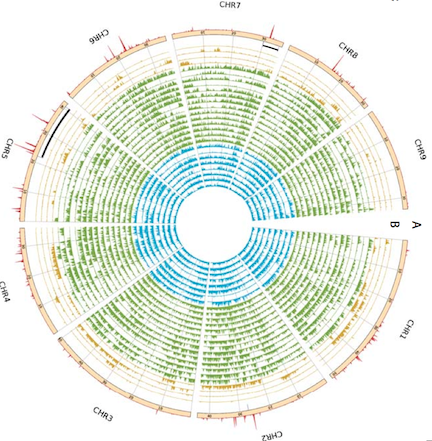
A high-quality assembly of the carrot (Daucus carota) genome is reported this week in Nature Genetics. Carrot is an important crop due to its high content of Vitamin A precursors, alpha- and beta-carotenes, as well as its popularity in global cuisines. The bright orange color of the modern carrot and its high carotenoid content are features that emerged through selection and breeding- the complete genome sequence will serve as a resource to aid breeders in crop improvement strategies.
Sequencing the carrot genome allowed for the identification of two novel Whole Genome Duplication events and 634 proposed pest and disease resistant genes. In addition, a novel candidate gene regulating carotenoid accumulation was found. Finally, the authors re-sequenced 35 carrot species and outgroups to determine genomic regions associated with domestication and estimated genetic diversity. Further phylogenomic comparisons with other plants clarified evolutionary divergence between carrot and tomato, grape and kiwifruit.
We spoke with lead author Philipp Simon to get some background on the research.
How did you end up working on carrots?
The position I am in focuses on carrot genetics and breeding. It became advertised soon after I completed my Ph.D. in genetics. The ability to do genetic research on a crop with a strong positive impact on consumers appealed to me. I was fortunate enough to enter that position.
What do you consider your most surprising result coming out of sequencing the whole genome?
The discovery of a candidate gene for the Y locus, which conditions the accumulation of carotenoid pigments in carrot roots. In previous work we were able to map the trait and also genes for enzymes in the carotenoid biosynthetic pathway, but none of those genes involved in carotenoid biosynthesis mapped with the Y locus. With a well-characterized genome available, we discovered a candidate for that important gene. The Y locus is one of the two genes responsible for the domestication of wild white carrots (ancestral wild type) to orange.
What user group do you think will benefit the most from these data?
The immediate users of the whole genome sequence will be by plant breeders for marker-assisted selection they have underway for carrot disease resistance and seed production traits. There are also several public sector labs doing more basic research on carrot pigments, biotic and abiotic stress response, reproduction, and evolution that will find it useful.
You propose an interesting model for carotenoid accumulation in the carrot. How might this knowledge be applied to the potential improvement of other crops?
There are several possibilities. The knowledge of this mutation in carrot may provide insights for identifying similar mutations in sequenced genomes of other crops, or generating similar mutations with genome editing technologies, for example. This could have application with other root crops such as cassava, but similar mutations are also known to influence pigment accumulation in fruit crops, so there may be applications beyond root crops.
What are some of your future directions going forward now that the genome assembly is complete?
Now we are using the carrot genome to understand genes for other carrot traits, including traits influencing accumulation of carotenoids, anthocyanins, carbohydrates and flavor terpenoids; pest and disease resistance; abiotic stress responses; plant reproduction and growth.
Bonus- do you have a favorite carrot recipe?
Regarding carrots in my diet, I usually eat raw carrots, but roasted or stir-fried carrots are also very tasty.