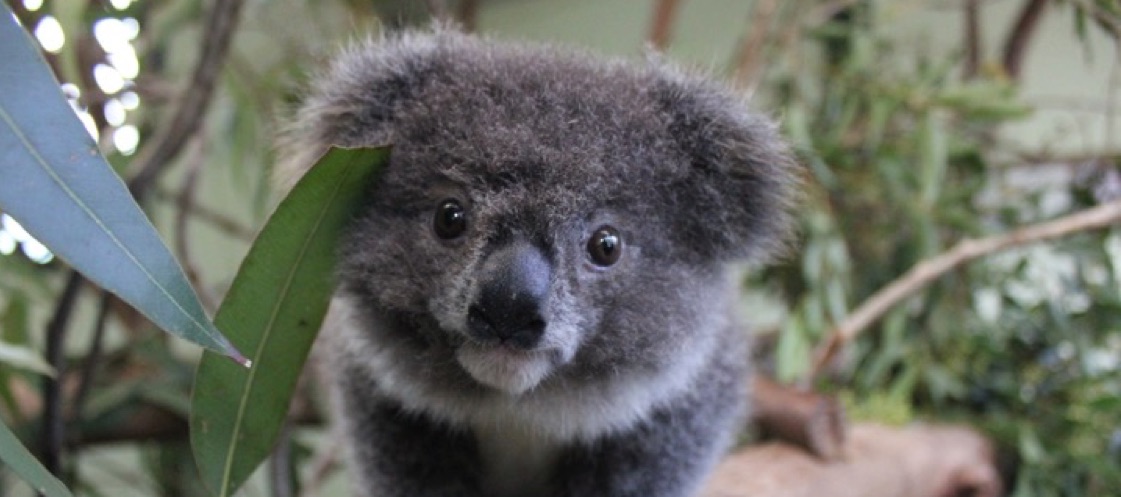
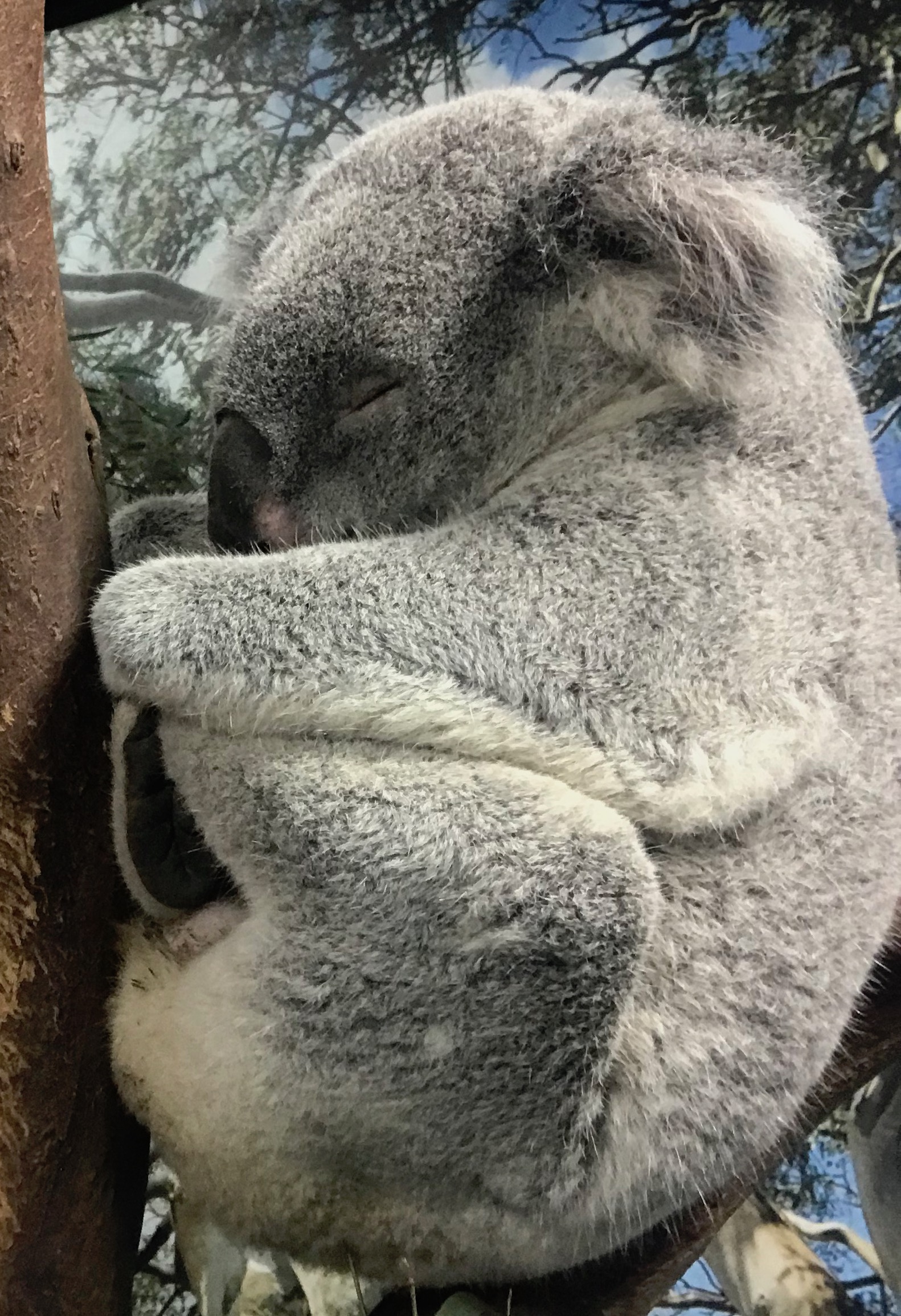
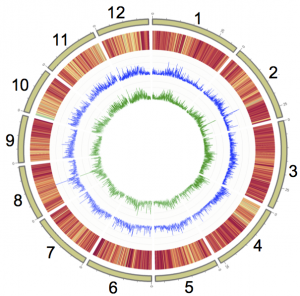
The genome assembly of the koala is reported in a paper published online in Nature Genetics. This high quality genome represents the most complete genome sequence for a marsupial to date. The data give insight into the highly specialized koala diet, consisting of eucalyptus leaves, and provide information that may be useful to combatting infectious disease.
Koalas are a vulnerable species and part of the aim of the the project was to use the genomic data to inform conservation efforts. We spoke with lead author Rebecca Johnson to get some background on this work:
How did the koala genome project come to be?
The genome project started as a small group of Australian researchers (from the Australian Museum, University of the Sunshine Coast and University of Sydney) who were enthusiastic about koala conservation and using genomics to manage populations and diseases. We partnered up with colleagues at the Ramaciotti Centre at the University of New South Wales (UNSW) who were enthusiastic to try out their new sequencing equipment on a ‘de novo mammal sized genome’. This hadn’t been done before in Australia.
We decided to take a bit of a risk and announce to the world in 2013 that we were establishing the Koala Genome Consortium and sequencing the genome. This was a very effective way of getting our project on the scientific horizon but then the pressure was on us to deliver! Fortunately for me (and the koala) one of my biggest career risks (announcing the genome well ahead of time) has resulted in a brilliant collaboration of scientists producing a high quality genome with many exciting outcomes and applications.
What do you think were the most interesting or surprising findings that came out of the genome data?
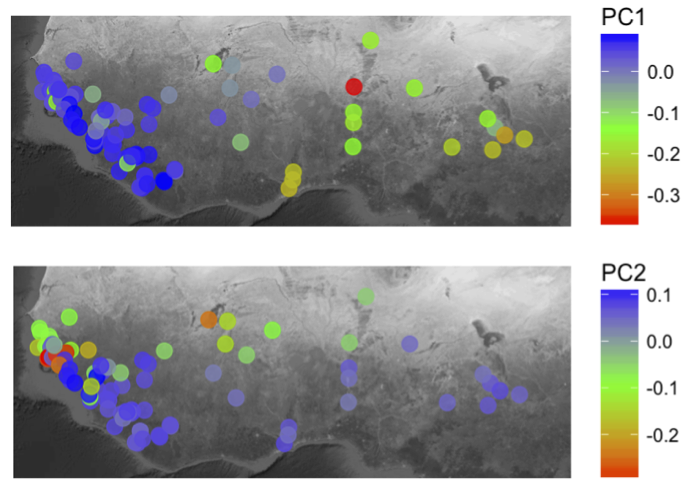
So many interesting things have come out of this work, so it is difficult for me to pinpoint one in particular. However, as a conservation geneticist I’m particularly fond of the conservation genomics work, particularly the historical population reconstruction which infers what koala populations would have looked like through evolutionary time. It was a little surprising to discover that koalas underwent such a dramatic decrease in population size 30-40kya, which was around the time many of the megafauna were experiencing extinction in Australia. Another surprise was that the three koalas used for this analysis are from two quite geographically separate locations (~600 km apart) but both suggest a dramatic reduction in population size indicative of widespread pressures across the continent.
Having this ‘deep-time’ perspective on koala populations, combined with the contemporary population work we did as part of this study we have a long term understanding of koalas in the landscape (i.e. the importance of long-term regional gene flow). Conservation management efforts can now be based on this holistic knowledge rather than a single genetic snapshot taken in time.
What are the biggest threat to the koalas now?
The koala is now classified as ‘vulnerable’ due to habitat loss and widespread disease. Threats to koalas are multifaceted, with the biggest primarily due to loss and fragmentation of habitat, urbanization, climate change and disease. Current estimates put the number of koalas in Australia at only 329,000 animals (range 144,000-605,000), and a continuing decline is predicted unless measures are put in place to arrest this decline.
How do you envision that this genomic information can aid conservation efforts?
The benefit of the genome to conservation efforts is widespread. The population diversity information presented in our work provides the impetus for a conservation management strategy to maintain gene flow regionally while incorporating the genetic legacy of biogeographic barriers. We have also identified the huge contrast in genome-wide levels of diversity across the northern and southern populations of koalas which will be factored into future decision making. The importance of genetic diversity indices for koala conservation has been included in the recently released NSW koala strategy so we will be focusing on highlighting the genetically healthy koala populations and ensuring they maintain regional gene flow. If more intensive measures such as translocations are required (for example from the genetically diverse populations to the genetically depauperate populations), we now have the tools and data to inform those decisions.
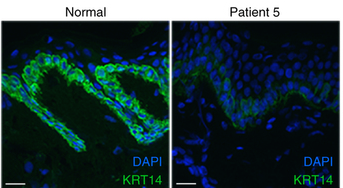
The immune gene repertoire we report as part of the genome is also being used directly in efforts to understand the response of koalas to disease such as chlamydia and the koala retrovirus (KoRV). Several of our collaborators on this work are involved in very important work developing and trialing vaccines for both chlamydia and KoRV. The genome affords the ability to understand which immune genes are up or down regulated in response to disease or treatment and provides the platform for future therapies to be tailored to the genome level.
What is it like working with koalas? Do you have any good stories that you would like to share?
It never gets tiring working with koalas and it was not difficult at all to bring collaborators on board to work on this project!
Koalas are notoriously chilled out animals (spending most of their time sleeping or eating), although my friends and colleagues who wrangle them in the field do report how unpleasant it is to be on the receiving end of their extremely sharp claws and nippy diprotodon teeth!
As part of sequencing the genome, our efforts to extract suitable quality DNA from koala blood were unsuccessful (possibly because they have a high lipid content in their blood) the only way we could get suitable quality DNA was to wait for an animal to be euthanized so we could access tissues suitable for genome and transcriptome work. Our two females were euthanized because they had advanced untreatable chlamydia. It is an extremely sobering experience to be involved in these necropsies because you can see the ravages of the disease on the body. While these moments are very tough they also inspire you to work harder to ensure we are producing the best possible science to conserve this amazing species.
For more video information, please see: