A new study published online this week in Nature Genetics reports the discovery of novel host targets of HIV infection identified from a high-throughput CRISPR/Cas9-based screen. This screen was performed in CD4 + T-cells and was designed to find candidate genes required for successful HIV infection, but whose inactivation did not affect cell viability. In this way, potential drug targets for anti-HIV therapy could be discovered.
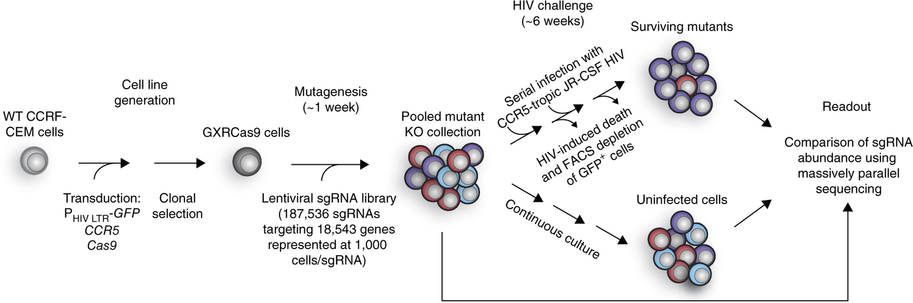
Park et al., Nature Genetics 2016
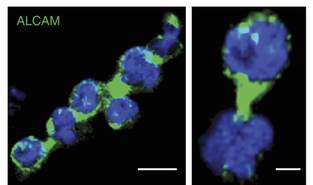
Park et al., Nature Genetics 2016
The authors found two known (CCR5 and CD4) and three novel (ALCAM, SLC35B2 and TPST2) cellular factors that, upon abrogation, prevented HIV infection but did not have any negative effects on the cell itself. These new genes are involved in sulfation and cell aggregation pathways and represent candidate targets for interventional HIV therapy.
We spoke with first author Ryan Park to get some background on this research:
Previous screens for host factors affecting HIV pathogenesis found a high number of hits, with low reproducibility across screens. With your CRISPR/Cas9 approach, were you expecting similar results? Did the low number of hits in your screen surprise you?
We designed our screen stringently, as the existing literature has not been clear on what genes would potentially serve as good targets for host-directed anti-HIV therapies. Our goal was thus to identify these host factors with high confidence while maintaining an unbiased approach. The very low number of hits was certainly surprising, though, as you note, the limited overlap among the previous screens raised the suspicion of a high false positive rate and/or low reproducibility.
You find three novel genes that are dispensable for cell viability but that are needed for successful HIV infection. Do you think that there could be natural polymorphisms in these genes in human populations that might mitigate susceptibility to HIV entry and transmission?
In the Exome Aggregation Consortium (ExAC) dataset recently published in Nature, there are individuals with truncations and/or homozygous mis-sense mutations in each of the three genes, as well as ITGAL (the loss of which we find is protective against HIV infection in primary CD4+ T cells). More work remains to be done to determine whether these individuals are relatively less susceptible to HIV infection.
Due to the high mutation rate of HIV and the emergence of resistance to drug therapies, potential targeting of host factors can be a useful strategy. Do you envision these findings being utilized to develop novel anti-HIV therapies?
Host-targeted HIV therapies are of great interest for multiple reasons. Firstly, as you note, the emergence of drug-resistant HIV strains remains a major issue, particularly in settings where adherence to a daily antiretroviral regimen is challenging. Drug-resistant strains are less likely to emerge in the face of incomplete adherence to host-targeted therapies. Secondly, the identification of host factors may also serve as a basis for gene therapies (in which gene editing is used to produce a population of HIV-resistant target cells) that could result in a permanent HIV cure. As noted above, more work remains to be done to determine whether inactivation of these genes protects against HIV infection at the organismal level without causing detrimental effects.
How might this screen be adapted to find host factors important at other stages of the HIV life cycle and do you have future plans to explore such work?
Our screen captured all but the latest stages of the HIV life cycle (particularly virion assembly, budding, and maturation); this is because HIV Tat, which drives the GFP reporter in our cell line model, is expressed prior to these steps. Development of an alternative reporter system that is activated by virion budding or maturation would allow identification of host factors involved only at these late stages. Because completion of the HIV life cycle is not required for host cell killing by HIV, cells lacking these late-acting host factors may still not be captured in a screen; more importantly, these late-acting host factors may therefore not be attractive therapeutic targets.
Can this screening method be employed to find host factors important for infection by other viruses? Do you speculate that there would be viruses for which a large number of non-essential host factors would be identified as important for infection?
The key elements of our approach, which include identification of a physiologically relevant cell line and the use of a high-complexity genome-wide sgRNA library, can be readily generalized to identify host factors that are critical to the propagation of any viral pathogen yet dispensable for cell viability. Our findings suggest that the number of non-essential host factors that are critical for HIV infection is quite limited, and that many candidate host factors identified by other screens or targeted studies may not be required for HIV infection or may compromise cell viability. Whether this is the case for other viruses is hard to know, but we have demonstrated that our approach can be quite powerful and specific in identifying the range of potential host targets with high confidence.