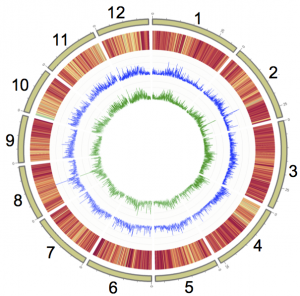
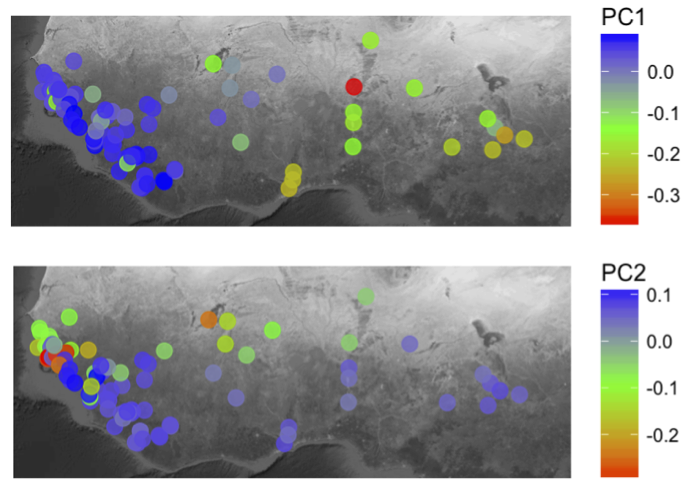
Rice is one of the most important crops on the planet, responsible for feeding billions of people. Given this global significance, studying rice in different geographies can be useful and aid in harnessing genetic diversity underlying particular traits and adaptations favorable to different environments. African rice (Oryza glaberrima Steud.) is mainly grown in sub-Saharan Africa and known for its stress tolerance. In a new article this week in Nature Genetics, Michael Purugganan and colleagues report the whole genome re-sequencing of 93 African rice landraces from various regions of Western coastal and sub-Saharan Africa. They create a genome-wide SNP map and through comparative genomic analysis study the domestication and population history of African rice. They use their map to perform GWAS for salt tolerance and find 11 significantly associated regions, highlighting the value of this unique genetic resource.
By studying various regions with distinct environments, the authors were able to get clues about adaptation and geographic spread of the populations. They focused on coastal Senegal and inland Togo, which have higher and lower levels of soil salinity, respectively, and interviewed farmers in the region to understand the agricultural practices they employ in each region. The knowledge of the farmers helped to inform the genetic analysis and contributed to the model of African rice domestication and dispersal.
You can watch some of the interviews with the farmers here:
African rice farmers- interviews
Additionally, we spoke with authors Michael Purugganan and Rachel Meyer to get some background on this research.
Why do you think that rice is understudied in Africa compared to other places?
MP: I think it’s because it is not widely grown, unlike its Asian counterpart which has pretty much taken over the world. But there definitely is more interest in African rice as breeders are trying to figure out how to increase food production in Africa, as well as to try to see what genes in African rice can be used to improve Asian rice.
RM: There is a lot of great research on improving Asian rice for African farmers that is being done by brilliant AfricaRice scientists, and they are working hard on the social science side too. But there are so many challenges that Africa disproportionately faces – particularly climate variation – that demands ramping up rice research. There is insufficient support for programs that integrate crop experiments and trials into the different farmlands. A better connection between scientists and small-scale farmers would really help farmers adopt new varieties too- because there is sometimes resistance to trying new ones.
How did you choose which samples to include in your analysis?
RM: Recognizing that a lot of NGO work encouraging farmers to grow Asian rice ramped up in the 80’s and 90’s, we took advantage of the germplasm largely donated in the 70’s to the West Africa Rice Development Association, which were duplicated and available through IRRI (International Rice Research Institute). We chose accessions with the most metadata available, preferring ones with georeferenced location and a cultivar name. It wasn’t until later that we realized water tables far inland were high in salinity, so we just tried to make sure we had a fair number of samples within 250km of the coast, or along rivers connecting to the ocean.
Were you surprised by any of your findings?
MP: There definitely were a few surprises in the data, but the big revelation for me was the long time for the population bottleneck that led to domestication. We found from the genomic data that it may have taken more than 10,000 years of steady population decline before full-blown domesticated African rice shows up in the archaeological record. This suggests the possibility that humans were already cultivating or managing its ancestor for thousands of years, and I think if this pattern holds for other domesticated crop species it will change our thinking on how domestication has taken place.
RM: I was surprised we got nice GWAS results with so few samples, and even more surprised that we saw several of those exhibiting signatures of geographic selection. We were lucky to find a broad distribution of traits in the landraces we chose to sequence, for we had made the DNA libraries ahead of the phenotyping experiments.
What was it like to meet and talk with the farmers?
RM: It was one of the highlights of my life to meet the farmers! I’m grateful to have gotten a glimpse of their heritage, their pride, and their struggles. We were all so impressed with the generosity of women, in particular, to help each other. We were also shocked by how many farms are run by the elderly; their children don’t see farming as profitable and many have left. For the three of us in the field, it made us think hard about how we can give back to the communities that gave us their time. I hope that crop science, publicity (like this blog) and policy changes can raise the profile of the small-scale farmer.
In each interview, the farmers also had a chance to interview us, and that part was especially interesting. Several asked really good questions about African and Asian rice domestication. You could see the cultural value of the basic science.
You chose to focus on salinity tolerance as a trait particularly relevant to farming in Africa. In what ways do you see your results being used for crop improvement?
RM: One of the authors, from AfricaRice, Dr. Kofi Bimpong, had actually been working on salt tolerance separately as well, and has two graduate student collecting African rice landraces in Casamance. If from this paper we can consider that domestication possibly occurred in the Inner Niger Delta region and also in the West, then these collecting efforts are all the more important because they are from a center of origin, promising more genetic variation than people would have ever estimated. If you look through the available germplasm there is so little that has been collected or studied from Casamance. It’s tricky collecting there, for there is social unrest, and landmines. Hats off to the young graduate students, Mamadou Sock and Bathe Diop, doing that fieldwork; I’m sure there is a lot of discovery to be made with those collections, and more promising salt tolerant landraces to integrate into breeding programs.
In addition, our results suggesting many of the salt tolerance genes are shared in both rice species make them more valuable to explore in other crops. Shared adaptive mechanisms are especially fascinating to evolutionary biologists and are powerful assets of the breeder’s toolbox.