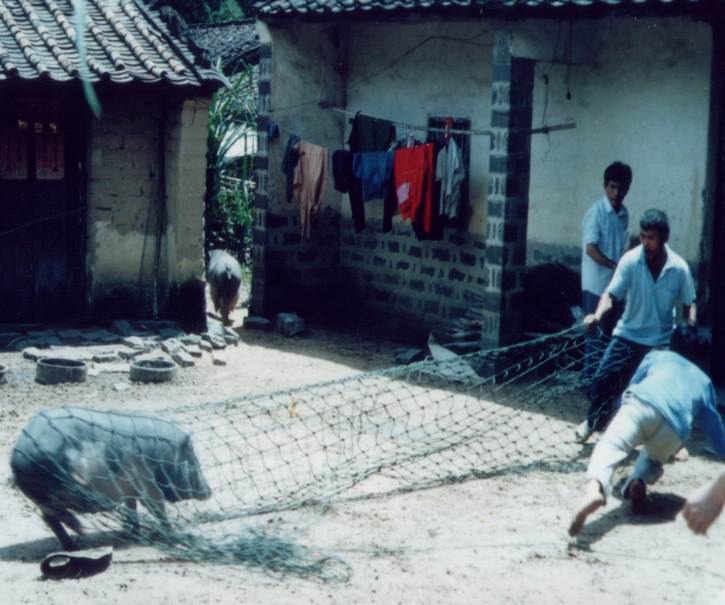
One species of wild tomato, Solanum lacerdae{credit}Sandy Knapp{/credit}
Most wild tomato species bear little resemblance to the large, red fruits you’re used to seeing in the supermarket. This is because humans have been molding the tomato to their own taste for thousands of years, by selecting for larger, tastier and (of course) redder fruits.
As a consequence of this selective breeding, we have significantly altered the tomato genome. A new paper published online this week in Nature Genetics analyzed the genomes of 360 tomato accessions, including multiple wild species and cultivated varieties, to understand exactly how and where humans have left their mark on the tomato genome.
This study, the product of a collaboration between many groups around the world, found that human selection on the tomato has led to vast improvement in certain traits at the cost of dramatically reducing genetic variation in large swaths of the genome. An unintended consequence of historical selective breeding in tomato is that there is now little room for improvement on many traits that we care about. By identifying these regions, the study will allow tomato breeders to make more strategic plans for future crop improvement.
We asked one of the study’s senior authors, Sanwen Huang, to tell us a little more about the work and why it is important:
This study was obviously a huge undertaking. How did collaborations come about, and what were the major difficulties in the project?
As an international consortium, we sequenced the tomato genome together (Nature 2012) and this project was regarded as another milestone of tomato research. The difficulty in the current project was deciding what to sequence. Fortunately, our team includes experts who understand tomato germplasm and they studied the natural variation of tomatoes for a long time. As a corollary, we combined tomato lines from many well studied core collections from several countries, such as the US (Roger Chetelat), Israel (Dani Zamir), France (Mathilde Causse), Italy (Andea Mazzucato), and China (Yongchen Du, Zhibiao Ye, and Jingfu Li).
What do you see as the most important aspect of your study’s results?
There are several important results that came out of this work. First, the evolution of tomato fruit size had two stages, from the wild progenitor of the modern cultivated tomato, Solanum pimpinellifolium, to cherry tomato (from ~1g to ~10g), and from cherry tomato to big-fruited tomato (from ~10g to ~100g). We found that there are two independent sets of QTLs or genes that have been selected during the two evolutionary stages. Second, there is a huge genomic signature of the divergence between fresh tomato and processing tomato [tomatoes used for commercial canning], on chromosome 5. This genomic region harbors several genes related to higher soluble solid content and fruit firmness that were selected during breeding for processing tomato. And more interestingly, we noticed that in recent fresh tomato F1 breeding, this region was also exploited for better taste and longer shelf-life. Third, we identified the causal variants for the pink tomato, which can be used for selective breeding. Pink tomato is a favorite in North China and I prefer it too, as it tastes better than the red ones. Finally, we found there have been costs to historical selection. For example, the near fixation of 25% of the tomato genome due genetic hitchhiking that occurred during domestication and improvement sweeps, as well as the linkage drags associated with wild introgression.

Cover of Nature, May 2012
Were you at all surprised to find such a large number of domestication and improvement sweeps? Did these results differ at all from other prominent vegetables, such as cucumber or potato?
The number and genomic proportion of domestication sweeps in tomato are similar to those in cucumber. However, the linkage disequilibrium blocks are bigger in tomato than in cucumber, possible due to the fact that tomato is a self-crossing species. Based on our data, we predict that the effective population size of tomato at domestication was about 300, similar to that of cucumber (~500), which is significantly smaller than that of maize (~150,000). This means these two vegetables have undergone much more severe bottlenecks during domestication as compared to maize.
How do you envision tomato breeders using the results of your study?
As a result of this work, tomato breeders will have a panoramic view of tomato variation and a better understanding of the raw materials used in their own breeding programs. From a practical standpoint, they will have access to a database of 11 million SNPs, from which they can pick the ones best suited to their molecular breeding programs. For example, they can combine the SNP dataset with their phenotypic data, to elucidate the genetic bases of important traits. Finally, and importantly I think, they will better understand the limitations of conventional breeding and the cost of historical selection, which will give them clues to improve their future programs.
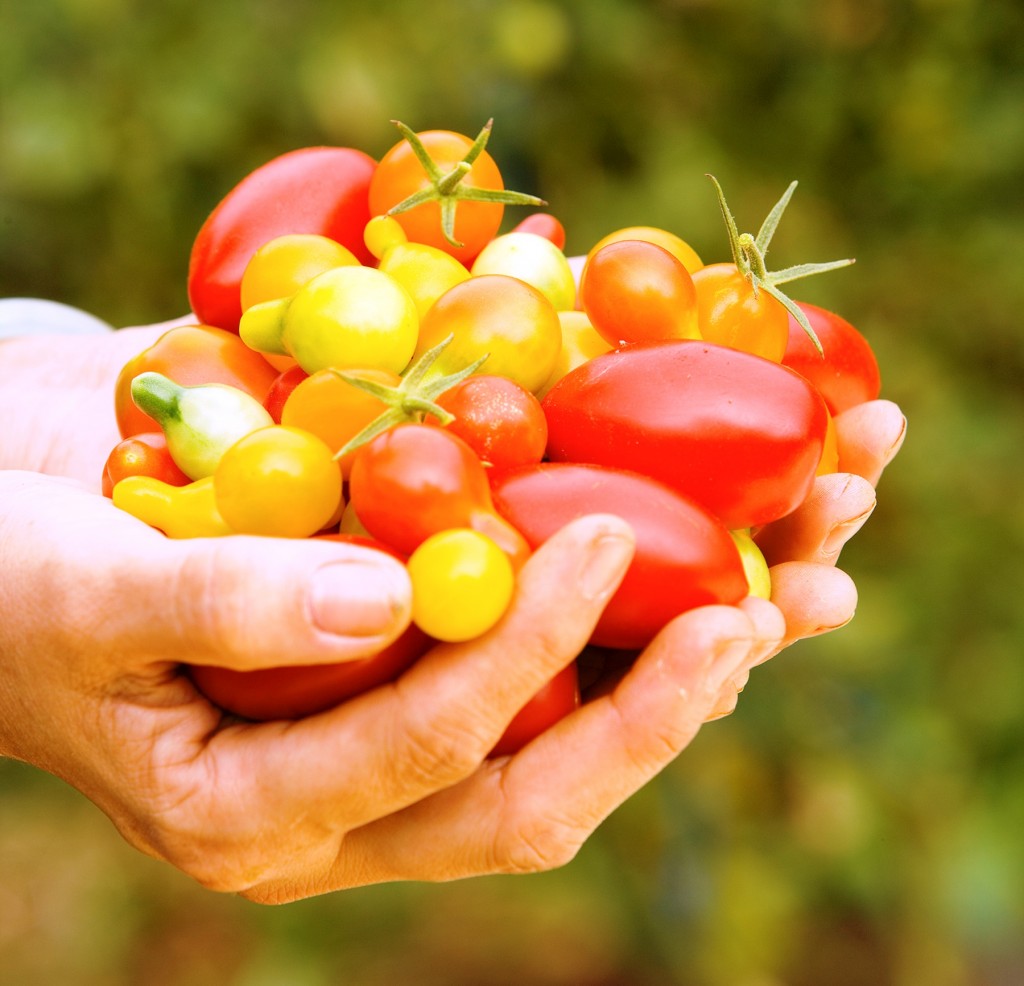

{credit}Photo courtesy of USDA Natural Resources Conservation Service{/credit}
Congratulations on your recent move to the Agricultural Genome Institute at Shenzhen where you are a co-founding director. Can you tell us a little about this new institute and what its goals are?
Thanks! The leadership of the Chinese Academy of Agricultural Sciences set up the institute (AGIS) to innovate agricultural research using genomics.
AGIS is located at the Dapeng District of Shenzhen, a beautiful bay area. The Shenzhen municipal government is developing the Dapeng Peninsula as the International Bio-valley and high-tech agriculture is one of the highlights. AGIS will recruit ~200 scientists who will decode, analyze, and utilize agricultural genomes. There will be three themes of research: the first theme is to develop basic algorithms and bioinformatic tools tailored for agricultural genomes, many of which are quite different from the human genome that has been the focus for most bioinformatians; the second theme is to empower agricultural breeding with genomics, to increase the efficiency and effectiveness of breeding that is essential to global food security; and the third theme is to provide genomic surveillance of food safety and agricultural environment, which is a huge concern of society and a need for sustainable development.

A vegetable market in Shanghai, China{credit}nadja robot via Flickr.com{/credit}
Bonus question: What is your favorite vegetable?
China is a country of vegetables, as there are over 200 kinds of vegetables that are regularly consumed in the country. I enjoy the diversity. For fruit vegetables I like tomato, cucumber, and chili; for leaf vegetables, I like Chinese cabbage, lettuce, and coriander.
You can read more about this exciting study at The Scientist. Read the full paper here.