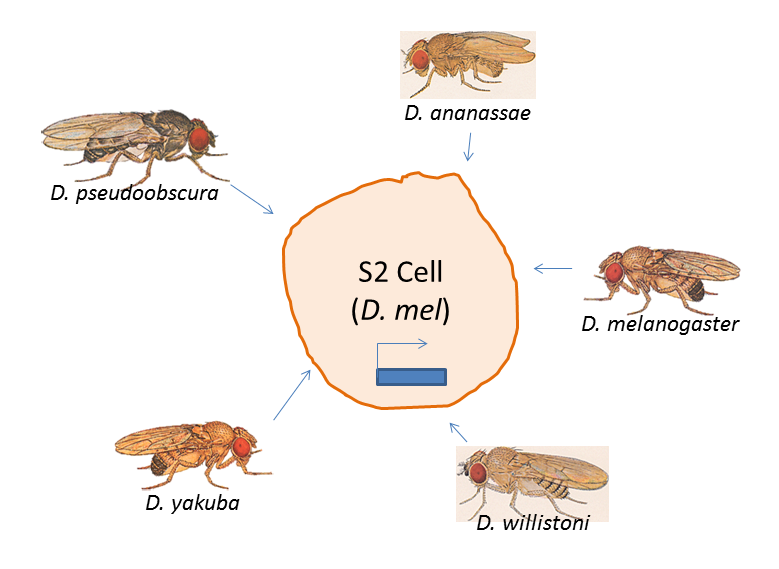
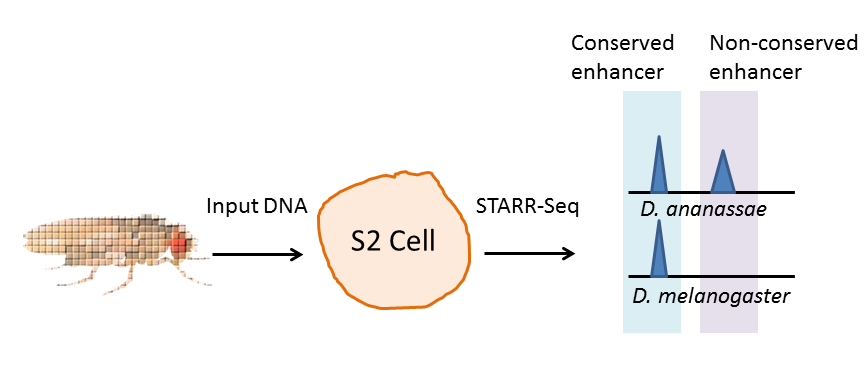
Two papers published online this week in Nature Genetics demonstrate that MYB, long known as a cancer gene, has many different strategies for driving tumorigenesis.
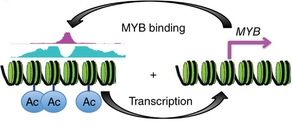
A positive feedback loop drives MYB overexpression in ACC{credit}Drier et al. Nat. Genet. 2016{/credit}
Bradley Bernstein, Birgit Knoechel and colleagues studied the role of MYB translocations in adenoid cystic carcinoma (ACC) and found that MYB translocations can reposition the gene to be driven by super-enhancers—which themselves are bound by MYB to drive its own expression even higher. In an interesting twist, they also found that MYB drives different regulatory programs in different ACC cell lineages: MYB’s oncogenic function is mediated by TP63 in myoepithelial cells, while in luminal epithelial cells, MYB appears to act through the Notch signaling pathway.
In an independent study focused on pediatric angiocentric gliomas, Keith Ligon, Rameen Beroukhim, Adam Resnick and colleagues found that MYB translocations resulting in MYB-QKI fusion genes are the most common MYB alteration in this cancer type. The fusion results in higher expression of MYB and loss of QKI expression, both of which contribute to the development of these gliomas. As in the ACC study, this translocation resulted in repositioning of MYB near enhancers that help drive its expression up. At the same time, the translocation caused loss of some regulatory elements, also leading to aberrant expression of MYB, and loss of function of QKI, a tumor suppressor. Thus, MYB-QKI uses three different mechanisms to drive gliomagenesis.
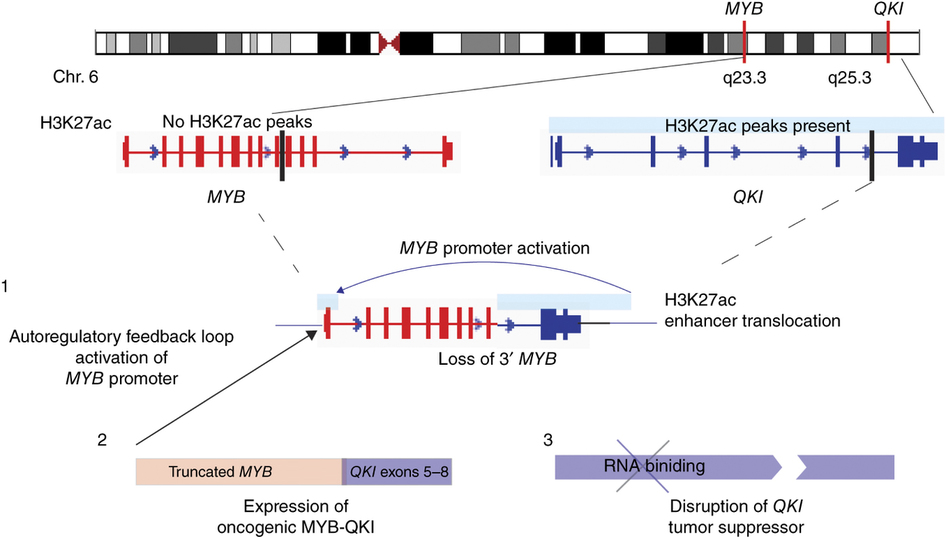
MYB-QKI promotes tumorigenesis through 3 mechanisms{credit}Bandopadhayay et al. Nat. Genet. 2016{/credit}
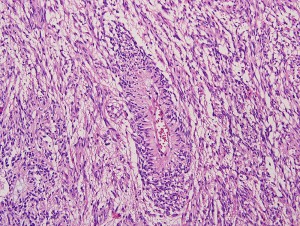
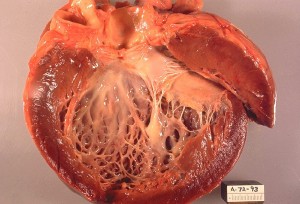
Angiocentric glioma. Angiocentric gliomas are characterized by cells that typically grow around blood vessels. {credit}Shakti Ramkissoon {/credit}
Both cancer types are relatively rare but aggressive, and new treatment options are sorely needed. Adenoid cystic carcinoma (ACC) occurs in secretory glands, mainly the salivary glands in the head and neck, and can spread to the nerves as well as metastasizing to distant sites, such as the lungs. The tumors are often resistant to therapy and can recur many years after the primary tumor has been removed surgically. Angiocentric gliomas are very rare brain tumors that generally affect children and young adults. Very little is known about the genetic changes that occur in this tumor type and, prior to this study, there were no known recurrent driver mutations, which are often good candidates for new targeted drug therapies. “The discovery of a recurrent rearrangement in angiocentric glioma provides a clinically relevant diagnostic marker, and insights into the biology that drives these tumors,” said Pratiti Bandopadhayay, one of the lead authors of the study.
We asked some of the authors from both studies to tell us a little more about the work and why it is important. Yotam Drier and Birgit Knoechel talked to us about the study in ACC. Pratiti Bandopadhayay, Lori Ramkissoon, Guillaume Bergthold and Payal Jain talked to us about the study in angiocentric gliomas.
How do your findings clarify earlier results showing a role for MYB in ACC? Do you think these findings are relevant for other cancer types?
Yotam Drier and Birgit Knoechel (Broad Institute):
Our work identified a unifying mechanism for MYB over-expression in ACC. Persson et al. suggested in 2009 that MYB over-expression occurs where the MYB 3′ untranslated region (UTR) is lost. However, in most cases of ACC the MYB 3′ UTR remains intact, and we now describe that in all cases of detected MYB rearrangements in this cancer–independent of whether the 3′ UTR is retained or lost–MYB is being driven by hijacking MYB bound super-enhancers, thus creating a positive feedback loop. This is complementary to the previous model, and we believe that in those cases where the MYB 3′ UTR is lost, both mechanisms would contribute to increased MYB expression.
We believe that similar rearrangements involving enhancer translocations may contribute to MYB overexpression in other cancer types. For example, our colleagues at Dana Farber simultaneously report a similar mechanism of MYB activation in angiocentric gliomas.
How do the mechanisms described in your paper compare to what is described in the related paper by Drier et al.?
Pratiti Bandopadhayay, Lori Ramkissoon and Guillaume Bergthold (Dana-Farber Cancer Institute) and Payal Jain (Children’s Hospital of Philadelphia):
We were excited to learn about the findings from the Bernstein group as their findings compliment ours, in a completely different tumor type. We found that angiocentric gliomas harbor rearrangements involving the MYB and QKI genes, while Dr. Bernstein’s team focused on adenoid cystic carcinomas, which frequently have similar MYB rearrangements. Both papers show that MYB rearrangements result in aberrant activation of the MYB promoter to drive expression of the oncogenic fusion proteins, and that these fusion proteins then participate in auto-regulatory feedback loops to drive their own expression.
From your perspective, what was the most unexpected finding in this study?
Yotam Drier and Birgit Knoechel:
We were surprised by our finding that MYB orchestrates 2 opposing epigenetic states—a TP63-dependent program in myoepithelial cells and a NOTCH-dependent program in luminal cells. Thus, overexpression of a single transcription factor can drive distinct epigenetic states that depend on the cellular context in which the overexpression occurs.
Pratiti Bandopadhayay, Lori Ramkissoon, Guillaume Bergthold and Payal Jain:
The unexpected result of our study that we find very exciting is that this one single driver rearrangement contributes to tumor growth through multiple mechanisms. MYB-QKI rearrangements simultaneously drive expression of a fusion protein that causes cells to grow faster and form tumors, it changes the regulatory landscape of the gene to promote expression of this protein and it simultaneously disrupts a tumor suppressor gene (QKI) that in turn also makes the cells divide faster. We feel that this finding is likely relevant to a number of other pediatric and adult cancers.
How does the fusion with QKI impact the function of the translocated MYB and do you think it is necessary for its role in driving gliomagenesis?
Pratiti Bandopadhayay, Lori Ramkissoon, Guillaume Bergthold and Payal Jain:
The rearrangement with QKI results in displacement of regulatory elements on QKI towards MYB and these elements help drive expression of MYB-QKI. In addition, it disrupts the function of QKI itself, which is a tumor suppressor gene. We feel that the association with QKI is important in angiocentric glioma since the rearrangement between MYB and QKI occurred with such high frequency in our study.
What are the additional steps needed before your findings can be implemented in the clinic?
Yotam Drier and Birgit Knoechel:
Interestingly, while BET inhibition can slow tumor growth in low grade ACCs, high grade ACCs often show genetic activation of NOTCH and are thus amenable to treatment with gamma secretase inhibitors or other NOTCH targeting therapies. It will be important to evaluate whether combining BET inhibition with NOTCH inhibition may show additional effects over BET inhibition alone. It is conceivable that by adding the NOTCH inhibitor one might preferentially target the luminal epithelial cells which are characterized by a NOTCH driven regulatory program. This will need to be tested further in preclinical models. Moreover, the fact that grade 3 tumors failed to respond to BET inhibition requires further preclinical analyses. Identifying mechanisms of BET inhibitor failure which are just entering clinical trials will be of utmost importance in order to predict which patients may benefit from these.
Pratiti Bandopadhayay, Lori Ramkissoon, Guillaume Bergthold and Payal Jain:
We are excited that our results provide us with novel possibilities to treat angiocentric gliomas. As MYB is a transcription factor the likelihood of targeting it or the MYB-QKI fusion is challenging; however we identified several downstream targets that represent potential therapeutic strategies. In addition, the finding of altered regulatory elements represents another exciting therapeutic strategy. Our findings directly impact clinical care for children with angiocentric glioma through development of two diagnostic tests that will be used to support the diagnosis of angiocentric glioma. We also feel our findings are likely relevant to other pediatric and adult cancers that are driven by driver rearrangements.
Finally we would like to highlight that multiple institutions and funding sources helped facilitate this study. We would also like to acknowledge the families whose children have been afflicted with Pediatric Low-Grade Glioma.
 This month’s cover image is inspired by the Article on p. 528 of this issue, by Jeff Wall, Nicola Illing, Nadav Ahituv and colleagues. The paper reports the genome of the bat Miniopterus natalensis and transcriptional dynamics in the developing bat wing. This species, one of a group known as vesper bats, is also known as the Natal long-fingered bat and is found in parts of Africa.
This month’s cover image is inspired by the Article on p. 528 of this issue, by Jeff Wall, Nicola Illing, Nadav Ahituv and colleagues. The paper reports the genome of the bat Miniopterus natalensis and transcriptional dynamics in the developing bat wing. This species, one of a group known as vesper bats, is also known as the Natal long-fingered bat and is found in parts of Africa.