This guest blog comes from Sowmya Swaminathan, Head of Editorial Policy and Research Integrity at Nature Research.
We are pleased to share results from a pilot with 13 journals that tested the Materials Design Analysis Reporting (MDAR) checklist, a minimum standards reporting checklist for the life sciences.
The MDAR framework, a minimum standards reporting framework for life sciences, was designed to provide a harmonizing principle for reporting requirements currently in use at various journals. It is meant to be flexible to adapt to various journal policies, and provides two levels of reporting stringency, a minimum recommendation and a best practice recommendation. The checklist was designed as an optional instrument to help adoption of this reporting framework. A statement of task can be found here.
We are very grateful to the 13 participating journals and platforms – BMC Microbiology, Ecology & Evolution, eLife, EMBO journals, Epigenetics, F1000R, Molecular Cancer Therapeutics, Microbiology Open, PeerJ, PLOS Biology, PNAS, Science, Scientific Reports – for testing the MDAR checklist.
The pilot had two main goals: first, to understand whether the checklist was accessible and useful for authors and editors to help comply with journal policy and second, to understand whether the elements within the checklist are clearly conveyed so as to help fulfil policy expectations. In total, 211 authors across participating journals tested the checklist and provided their feedback. Participating journal teams screened 289 manuscripts using the checklist and 89 of these manuscripts were subject to a dual-assessment by independent reviewers, which allowed us to determine inter-assessor agreement, and thus clarity of specific items on the checklist.
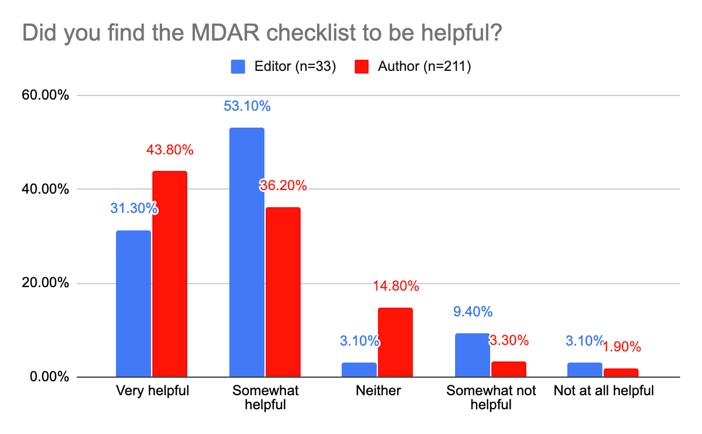
We are encouraged to find that about 80% of authors and editors found the checklist useful to different degrees and that the majority of participating editors reported only a small increase in manuscript processing time as a result of using the checklist. While participating authors and editors did not identify major gaps in the requirements covered in the checklist, the feedback from authors and editors and the inter-assessor agreement results have given us a better understanding of areas in the checklist and elaboration document where the language is unclear and needs to be improved.
We are making the draft MDAR Framework, MDAR Checklist and MDAR Elaboration document and the pilot datasets available here. This work was also recently presented at the NASEM workshop on Enhancing Scientific Reproducibility through Transparent Reporting (slides available here).
We are currently gathering feedback on the MDAR framework, checklist and elaboration document from a broad group of about 40-50 experts on transparency and reproducibility. Based on the feedback from the pilot and the expert input, we anticipate revising all three MDAR outputs by the end of 2019.
We are sharing this update on the work of the Minimum Standards Working Group through coordinated posts on member platforms. If you would like more information about our work and progress, please contact Veronique Kiermer and Sowmya Swaminathan.
On behalf of the “minimal standards” working group:
Karen Chambers (Wiley)
Andy Collings (eLife)
Chris Graf (Wiley)
Veronique Kiermer (Public Library of Science; vkiermer@plos.org)
David Mellor (Center for Open Science)
Malcolm Macleod (University of Edinburgh)
Sowmya Swaminathan (Nature Research/Springer Nature; s.swaminathan@us.nature.com)
Deborah Sweet (Cell Press/Elsevier)
Valda Vinson (Science/AAAS)