 Fluorescence microscopy has transformed the life sciences. By attaching fluorescent dyes or proteins to cellular structures, researchers can image fine cellular morphology; track molecular localization, motion, and dynamics; and more. But fluorescence microscopy also presents significant obstacles. One of those is multiplexing.
Fluorescence microscopy has transformed the life sciences. By attaching fluorescent dyes or proteins to cellular structures, researchers can image fine cellular morphology; track molecular localization, motion, and dynamics; and more. But fluorescence microscopy also presents significant obstacles. One of those is multiplexing.
Tag Archives: imaging
New neuroscience tools for team science in ‘big data’ era
By Esther Landhuis
Wandering the convention center among 30,000-plus researchers, students and vendors at the Society for Neuroscience annual meeting in San Diego last November, I struggled to wrap my head around a feature I was writing for this week’s Nature, on managing big brain data. Mice, molecular biology and cell sorting reigned supreme in my former life as a bench scientist. Neurons, brain imaging, terabytes — not so much. So when it came time to find an entry into the vast universe of the brain, I latched onto something that seemed small and manageable: the fruit fly.
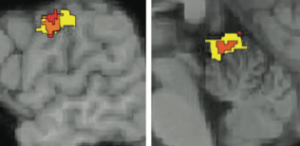
Ann-Shyn Chiang of National Tsing Hua University, Taiwan, told the SFN crowd his team has spent a decade imaging 60,000 neurons in the Drosophila brain. The pictures produced 3D maps detailed enough to show which neurons control precise behaviors, such as shaking the head side to side (see video). But here’s the part that blew my mind: They aren’t even halfway done (flies have 135,000 brain neurons), and mapping the human brain with similar methods would take 17 million years!
You are currently viewing a placeholder content from YouTube. To access the actual content, click the button below. Please note that doing so will share data with third-party providers.
Head shake behavior elicited by a 593.5-nm laser. Credit Po-Yen Hsiao and Ann-Shyn Chiang.
‘Google Earth’ of the brain slated for planetarium show
If you’re anything like me, you love a good planetarium show. I don’t mean the trippy laser light shows set to Pink Floyd tunes (although these certainly have their place), but rather the kind of immersive experience that gives you a glimpse into the untold depths of the universe and a few wondrous moments of what it feels like to soar through outer space. Now, a team of neuroscientists, astronomers, software engineers and film specialists are working on a new planetarium show to give us a fly-through experience in a different kind of vast and awe-inspiring space: the human brain.
You are currently viewing a placeholder content from Default. To access the actual content, click the button below. Please note that doing so will share data with third-party providers.
The project is called the Neurodome. It’s the brainchild of Jonathan Fisher, a neuroscientist at New York Medical College who also has a background in astrophysics. Although the planetarium version of the Neurodome is not yet complete, you can get a taste of what to expect tomorrow evening at Columbia University in New York, where Fisher and Columbia astronomer Matt Turk will guide viewers through pictures of outer space and images of human brains, explaining how light travels across the universe from distant stars and into the eye, triggering electrical impulses in the brain’s neural pathways. “Just like you can walk through Google Earth, we’ll walk through the brain,” says Fisher.
Call and response
The (highly abbreviated) life story of a paper appearing in Nature often goes something like this: ideas are birthed and experiments envisioned. Pilot experiments are run, yielding beautiful preliminary data. Replication and controls are then gathered over the course of months, if not years of hard labor. The paper is written, submitted, and reviewed. A few (two is typical) rounds of review and revision later, it is published (with highly variable degrees of reviewer and editorial unanimity). But this is by no means the end, rather, just a milestone in the evaluation process by the community. In journals, post-publication evaluation has traditionally occurred in the form of peer-reviewed follow-up papers or formal commentary. This may change someday as alternative forms of scientific publishing are explored, but for today we’ll talk about a formal addendum we’re publishing on a 2011 paper by Cathy Price and colleagues and invite you to add to the discussion.