by Myles Axton
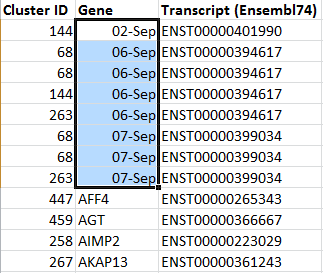
Normally we do not re-examine supplementary information in this detail, but there is a common minor problem that systematically affects a small number of gene IDs within long lists of gene names copied into spreadsheets in the supplementary tables of many articles. We suggest checking for this problem before submitting tables to journals. It is easy to see the altered gene names by sorting the column in a separate version of the file and then searching for the misspelled name to correct it in the replacement version intended for publication.
The authors of this paper claim that gene names in a large proportion of papers reporting gene expression data have this problem. Here we list the supplementary tables they identified in the journal prior to 2015 and from the first nine months of 2016 that we found to contain one or more misprints. We think that these mistakes do not prevent reuse of the datasets provided and as stated in the accompanying editorial Legible ledgers we do not propose to publish formal Corrigenda for the supplementary tables of these articles.