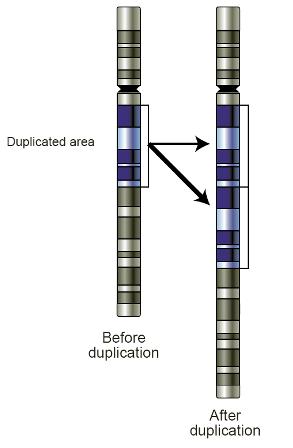
During the past 25 years, Nature Genetics has been lucky to publish many exciting papers, more than a few of which can be described as “landmark” papers—publications that have had a dramatic and long-lasting impact on a field. In 2004, the Journal published such a study by Stephen Scherer, Charles Lee and colleagues (Iafrate et al.) in which they reported 255 loci across the human genome containing large structural variants.
In 2017, the idea that there exist large numbers of structural variants in the genome (such as rearrangements, deletions and insertions) that differ from person to person is an established fact. But in 2004, this was not the prevailing wisdom. Prof. Scherer has already written an excellent essay at The Winnower about the study and its importance to the field, so I won’t recap it in detail here—I will simply encourage you to the read the piece.
Charles Lee wrote us about the study by email. “I saw a talk by Dr. Dan Pinkel at the 2002 ASHG meeting where he presented his latest array CGH findings,” he remembers. “In his talk, one of the slides showed the array CGH results of a trisomy 18 patient and Dan remarked how cleanly his array platform performed, especially for the other chromosomes. But in fact, I (and others, I’m sure) could see that there were actually occasional clones that deviated from the expected log2 ratio of 0. During the question period, I sheepishly asked him about these clones. I really didn’t mean to criticize his platform, but I think that he took it that way. Those “blips” bothered me and when I returned to Boston, John Iafrate (who was a postdoc with me at the time) began our own array CGH experiments. Ironically, there were several other groups that were way ahead of us with respect to technical expertise and experience with array CGH, but it could be that they considered these “blips” as technical artifacts – without biological implications.”
Prof. Lee added, “In late 2003, I gave a talk at the University of Toronto and met Stephen Scherer in person for the first time. In a casual conversation, we realized that we were both using the same 1 MB chromosome microarray platform from Spectral Genomics and that we were both seeing these recurrent ‘blips’ in our data.”
Stephen Scherer also corresponded with us by email about the study and the mutual decision to collaborate with the Lee lab. “We were both were fresh enough to look beyond what others were calling ‘noise’ to realize these aberrations represented intermediate and gene-level copy number variation.”
“Many of us suspected it was there,” he said of the large-scale variation they uncovered, “based on the fact there were lots of smaller indels and that 0.6% of the population carried cytogenetic alterations. We kind of predicted it in our chromosome 7 mapping and sequence paper, but only at the chromosomal level.”
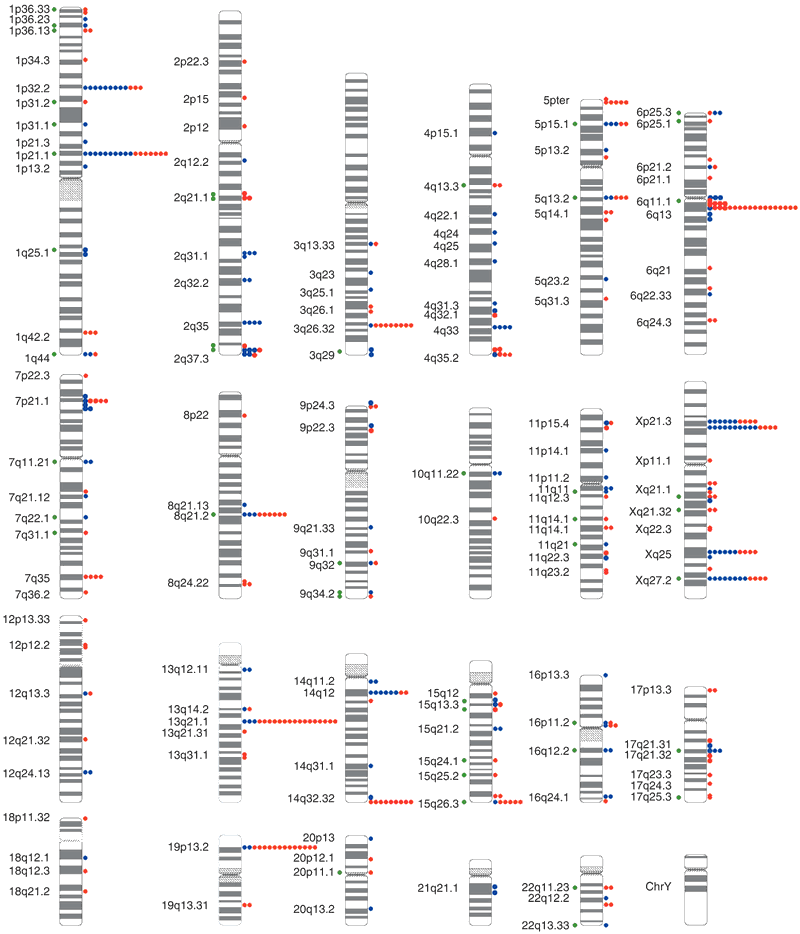
Circles to the right of each chromosome ideogram show the number of individuals with copy gains (blue) and losses (red) for each clone among 39 unrelated, healthy control individuals. Green circles to the left indicate known genome sequence gaps within 100 kb of the clone, or segmental duplications known to overlap the clone, as compared to the Human Recent Segmental Duplication Browser. Cytogenetic band positions are shown to the left. {credit}Fig. 1 from Iafrate et al. 2004{/credit}
The study by Iafrate et al. was published on August 1, 2004. Exactly one week prior, a very similar study by Michael Wigler and colleagues (Sebat et al.) was published in Science. The methods used by the two groups were different, but the findings and implications were consistent with each other. “Charles and I were happy to see the Wigler paper,” said Prof. Scherer, “because nobody believed our results.” Prof. Lee added, “This was one of the most difficult papers for me to publish. The reviewers were very skeptical. We had to keep providing more and more validation data, and one of the reviewers even commented that s/he did not believe that the paper was worthy of being an article and we had to shortened the paper into a Brief Communication. At the end, Reviewer #2, who was persistently negative wrote: ‘… I still feel hesitant about publication of this work in Nature Genetics… and I still doubt the importance and novelty of their work.” Prof. Scherer remembers similar levels of skepticism in the community. “Prior to publication I was showing the data at talks, including one at Michigan where they were trying to recruit me, and I remember getting trashed. People in my own department were mostly the same.”
[I looked up the referee reports and internal notes from the review process and Prof. Lee is correct that at least one of the reviewers was very skeptical about the impact of the study. However, I do want to note the very unusual fact, at least by today’s standards, that the study was published a little more than 2 months after initial submission, according to our records. I wish this was more common!]
After publication, however, the importance of the studies was immediately clear, at least to those working most closely in the field. Nigel Carter contributed a News and Views article in Nature Genetics about the studies. He wrote, “This unexpected level of LCV [large-scale copy-number variation] forces us to re-evaluate our view of the structure of the normal human genome.”
However, Prof. Lee remembers some ongoing skepticism about the work. “For more than 18 months after the paper was published, I had trouble getting grant funding for continuing my work in human copy number variation. Some comments that I received included, ‘If this was real, the Human Genome Project would have found it.’ I am embarrassed to say that I was forced to write for smaller grants on other topics and when funded, did everything I could to complete the projects using less money and use the ‘extra’ funds for my human copy number variation interests. It was very, very frustrating.”
In 2007, Science announced Human Genetic Variation as the Breakthrough of the Year. “When I saw this article in Science,” Prof. Lee said, “I felt like there was finally some widespread acceptance of our findings in the general scientific community.”
“However, this came with different issues.” For example, he often received the response from the GWAS community that structural variation is interesting, but it is too difficult to incorporate into GWAS. “So, most association studies continued to focus on SNPs, which is a problem that persists to this very day.”
The findings in Iafrate et al. were based on, by today’s standards, a fairly small sample of 55 individuals profiled by array comparative hybridization array comprising ~12% of the genome (the study in Science reported results from 20 individuals using representational oligonucleotide microarray analysis). However, the impact on the field was anything but small. Part of the legacy of the studies was the establishment of the Database of Genomic Variants (originally the Genome Variation Database) that has now collected over 550,000 CNVs. The discovery that so many structural variants are present in our genomes, even in healthy individuals, opened up an entire field of study to understand the function of these variants, and much is still to be discovered (see for example a recent study on the impact of structural variation on human gene expression).
Prof. Scherer summed up the impact of the studies this way: “If you remember the fights between the public Human Genome Project and Celera Genomics, and them finger-pointing to the errors in each other’s assemblies, in many cases these were due to CNV and other structural variations. They had no idea these CNV variants existed. It was really the 2004 Nature Genetics and Science papers, coincident, pure discovery, that opened the eyes of the community and it took some longer than others to believe it.”