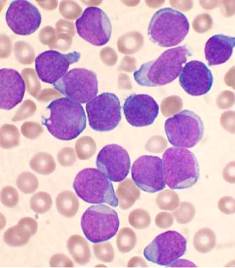
A Wright’s stained bone marrow aspirate smear of patient with precursor B-cell acute lymphoblastic leukemia. {credit}VashiDonsk via Wikipedia{/credit}
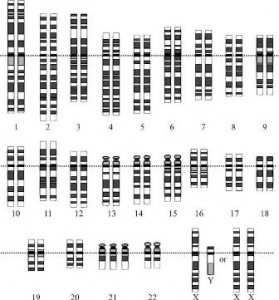
Trisomy 21 (having 3 copies of chromosome 21) is most well known as the cause of Down syndrome. But as you can imagine, having an entire extra copy of a chromosome has other negative consequences as well. For one, people with Down syndrome are 20 times more likely than the average person to develop a severe form of leukemia, B cell acute lymphoblastic leukemia (B-ALL). Two recent studies have helped further our understanding of the molecular disturbances that take place in trisomy 21.
In Nature, Audrey Letourneau et al. took advantage of a rare situation to identify the genes that are misregulated in Down syndrome. The researchers profiled the transcriptomes of identical twins that differed in one crucial aspect: one twin had 3 copies of chromosome 21, while the other had a normal complement of chromosomes. (The samples were collected post-mortem from the fetuses, with the permission of the parents). This approach allowed the researchers to avoid any noise from irrelevant differences, since the genes of both twins would be identical. (Read the article in The Scientist about this study here).
Not surprisingly, they found that trisomy 21 causes gene regulation problems on all chromosomes. Misregulated genes are organized along the chromosomes in domains, and these domains were defined by changes to the chromatin methylation patterns. Importantly for future research efforts, they also showed that the corresponding genomic regions in the mouse model for trisomy 21 were similarly modified compared to control mice.
In a second paper published online this week by Andrew Lane et al. in Nature Genetics looked specifically at the relationship between Down syndrome and B-ALL. The authors identified two genomic events as the drivers behind Down syndrome-related B cell acute lymphoblastic leukemia (B-ALL): overexpression of the nucleosome remodeling protein HMGN1 and changes in histone methylation marks. (You can read the Dana-Farber Cancer Institute’s press release about the study here).
Through a very meticulous set of experiments, they first show that just having an extra copy of a small region of chromosome 21 (or in this case, the corresponding mouse chromosome, 16) with 31 genes is sufficient for giving B cell precursors the ability to self renew indefinitely—the first step to cancer formation. From there, they identify and confirm a single driver gene on chromosome 21, HMGN1, as being expressed at unusually high levels. This high expression of HMGN1 causes a decrease in one type of methylation (H3K27me3), leading to overexpression of genes usually carrying both H3K27me3 and another histone mark, H3K4me3.
Interestingly, the authors of Letourneau et al. mention HMGN1 as a good candidate for regulating the genome-wide chromatin modifications they found. The accompanying News & Views article by Benjamin Pope and David Gilbert note that HMNG1 should be a target of future study in Down syndrome. Looks like the authors of Lane et al. got the message far in advance!
So a pattern emerges: changes in chromatin methylation patterns are a key event in trisomy 21 overall and in Down syndrome-associated B-ALL specifically. Dr. Lane, lead author of the paper in Nature Genetics wonders “Could this [chromatin modifications] be a unifying theme for phenotypes (not only cancer) associated with DS?” As I mentioned earlier, having 3 copies of this chromosome is bad for a number of reasons: higher risk of B-ALL and testicular cancer, vision and hearing problems, thyroid issues, higher risk of type I diabetes, gastrointestinal issues, low or no fertility and the more widely known neurocognitive isssues. Future studies on epigenetic changes in Down syndrome, and the regualtion of HMNG1, should be able to unravel the mechanisms underlying these different aspects of Down syndrome.
Outside of Down syndrome, these 2 studies may also lead to a better understanding of (and hopefully new treatments for) cancers caused by epigenetic changes. As the article by Lane et al. showed, changes to the chromatin landscape allowed B cell precursors to make that first step toward leukemia. By understanding how this happens, we can start to find ways to prevent it.