Q&A with Ronald Walsworth who is a staff scientist at the Harvard-Smithsonian Center for Astrophysics and a faculty member in the Harvard physics department.
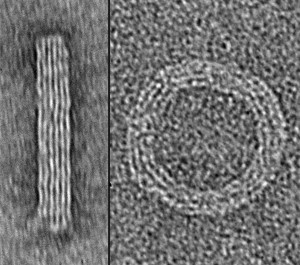
The scientist profiled in the August issue of Nature Methods, Ronald Walsworth, who is a physicist at the Harvard Center for Astrophysics (CfA) has built and tested a quantum diamond microscope that benefits from particular flaws in a diamond.
What follows is an edited excerpt of his conversation with Nature Methods’ technology editor Vivien Marx. Read more here.

Ronald Walsworth (r) and Chih-Hao Li (l) adjust a laser frequency comb used in the search for Earth-like exoplanets.
Photo credit: Harvard-Smithsonian Center for Astrophysics
Q: The new instrument can quantify single cells, what else can it or might it do?
RW: We have all these neat, cool things that diamond sensing can do, based on the way it helps to detect small changes in magnetic fields. Diamonds can go into extreme environments–underground, under water, or they can go into high temperatures such as in an airplane engine–where there is a need for sensors.
Some companies are developing imaging systems based on our research on the special kind of flaw in diamonds that involve nitrogen vacancy (NV) centers. There is an entire community of researchers working on NV centers who use them as nanoscale probes with which you can map out magnetic signatures at the near-atomic scale.
There are physical science applications such as nano-scale probing of surfaces of novel materials that might be used in computing or for energy storage. They are also used for condensed matter physics research. In the life sciences, these diamonds can probe living tissue. We’re pursuing an approach using a planar surface with many NV centers to image a sample. In our paper, we showed single-cell imaging and we think we could move toward single molecule sensing and imaging, such as assaying tissues for magnetic signatures in early-stage brain disease.
Q: You do fundamental physics. Isn’t biology too squishy for you?
RW: We are doing a lot of basic physics research, a bit of astrophysics, lots of different things. I enjoy learning new things. One way to do that and to be professionally productive is to develop new tools that are relevant from day one for some field that is new to me. Then I have the enjoyment of learning while I am contributing and while I’m still kind of ignorant. About half way up the learning curve is where I often have my best ideas.
I am involved in collaborations about how to create networks of atomic clocks, another is about sensing gravity waves, or new ways for detecting exoplanets. I am a professor in the department of physics and a member of the center for brain science. I have a lab there, too, next to [neuroscientist] Jeff Lichtman. Some astro-people are increasingly interested in aspects of biology, too. There’s the Origins of Life Initiative at Harvard led by my friend and exoplanet astronomer Dmitar Sasselov, who is also interested in questions of synthetic biology.
Q: Your background led you down this path?
RW: I did my PhD in physics at Harvard and kind of never left. I never really did a post-doc. I got my PhD in the physics department on atomic clocks and fundamental symmetry tests in physics and did some work at the CfA where there was an atomic clock group. Then I got an offer to do a post-doc with future Nobel Laureate David Wineland, at the National Institute of Standards and Technology. I felt that was great, but I wanted to finish up my PhD work.
One thing led to another, and after a year I developed some ideas of my own that I wanted to pursue. There was some unused lab space and I took a gamble that I could just keep myself going on my own and do the research that I wanted to do and declined the post-doc. Years passed and I was able to raise money and build up a research group from scratch.
Q: You said you don’t recommend this path to others, why might it not work for them?
RW: The Smithsonian had some funding that made me a principal investigator day one. You didn’t have to go through some formal hiring process, you could just try things. If you got resources to keep yourself afloat, you did science. It was very fertile and it gave myself and others a path through which to rise up, if we could get science done.
By the mid-1990s, I had built a group of four to five people, without being hired by anyone in any formal way other than having people at CfA say: “you can stick around in some space in a corner.”
I never went through a formal hiring process following up on an advertised position. I’ve never been hired anywhere. The last time I applied for a post was in 1984 when I applied to graduate school.
Q: Is that kind of path still possible?
RW: There is a good aspect to having more formalized procedures, there isn‘t nepotism and favoritism. Under-represented groups can be properly advanced and you are making sure there is a level playing field: those are all good things.
When it’s a bit more like the Wild West, there are lowered hurdles for young scientists when they are creative and full of energy. But even when you are trying to make things progressive and formalized it becomes unintentionally regressive to satisfy older and middle-aged scientists on hiring panels and study sections who vet and decide that “this is the young person we are allowing to join our ranks.” That lets existing researchers decide who should be hired and given a chance, rather than giving everybody a chance to pursue their creativity and see how it works out.
Q: How do you personally advise young scientists?
RW: Now I am going to sound like a griping middle-aged guy but I can think back to the young guy, too. There are too many barriers keeping people from trying things out. Scientists write proposals to get funding. We are enthusiastic about some of the proposals. Others we are less excited about. It’s tough when those ones are the ones we are less excited about.
But we always need to look out for the younger people, the next generation of scientists with promise and talent. We need to clear the path so they can get things done. I have physics graduate students from MIT and Harvard, some are also in chemistry or biophysics, some are MD/PhDs. I try to integrate everybody. They are in different locations, so we switch locations for lab group meetings. Fridays we have group seminars and then we go out to lunch together. One of my post-docs is someone who should already have a job. That’s another thing that bothers me, there are not enough jobs.
There are jobs for people who want to leave science who use their great analytical skills to do data analysis in the financial world. But for people to deploy their skills properly and continue in science, it’s hard. I have helped land good jobs for some of my people and I have a few more who are just finishing post-docs, who are just great and who need jobs.