Tuning reagents, software, or equipment is all in a day’s work in the lab. Building instruments from scratch, however, is a task more typical for physicists who might 3D print or machine the parts they need and then assemble them into the instrument they want. They might construct an instrument for a specific experiment or develop a design that helps hundreds of labs. That model could go on to be modified and hacked in a variety of ways.

In light-sheet microscopy, a sample is illuminated with a thin sheet of light and fluorescence is detected by a separate lens placed orthogonally to the excitation light. {credit}Vineeth Surendranath{/credit}

Build your own OpenSPIM system{credit}Michael Weber, Peter Pitrone, Pavel Tomancak{/credit}
In microscopy, biologists as well as physicists and computer scientists are building the hardware and software they want and sharing the blue-prints with others.
Building an OpenSPIM model is not quite this fast, but this shows the parts needed for those who want to give it a try.
Here are some user experiences from the OpenSPIM community. You can read more in the December issue of Nature Methods.
Perspectives from users, builders and one-day-maybe OpenSPIM builders

From left to right: Tiago Pinheiro, Johanna Gassler, Radoslav Aleksandrov, Florian Vollrath et al.
An OpenSPIM community has evolved to address the needs of researchers setting out to build their own systems.
Johanna Gassler, Tiago Pinheiro, Florian Vollrath worked together during the European Molecular Biology Organization (EMBO) course on light-sheet microscopy in August. Separately, Johannes Girstmair at University College London built an OpenSPIM microscope.
Johanna Gassler, Tiago Pinheiro, Florian Vollrath and Radoslav Aleksandrov worked together during the European Molecular Biology Organization (EMBO) course on light-sheet microscopy in August. Separately, Johannes Girstmair at University College London built an OpenSPIM microscope.

Johanna Gassler
PhD student in the lab of
Kikue Tachibana-Konwalski
Institute of Molecular Biotechnology
of the Austrian Academy of Sciences
Vienna, Austria {credit}Philippe Laissue{/credit}
Her thoughts on using light-sheet microscopy…
Gassler works with mouse oocytes and early embryos in a lab that looks at many facets of how an oocyte transforms into a zygote after fertilization. She does live-cell imaging with confocal microscopy and phototoxicity is a constant concern. Light-sheet microscopy lets her take a closer look, especially in terms of temporal resolution, at the dynamic processes inside an egg or an early embryo without having the types of phototoxicity worries she would have with other forms of microscopy.
In her view, light-sheet microscopy is one of the most exciting technologies of the last decade, “and it is really great to be a scientist in a time where these systems are still in their developing stage and to see how fast progress is made. “
“When imaging samples with confocal microscopy one does not tend to think so much about the specific characteristics of your sample compared to your neighbors. You just use the same microscope to image both, of course imaging settings change, but the hardware doesn’t. When taking the route of building your own microscope like with OpenSPIM, one is way more flexible in what pieces of hardware you would like to add to improve the imaging of your sample specifically. This flexibility is a huge advantage of OpenSPIM, but also a disadvantage at the same time. “
“If an OpenSPIM is built for a special application and the group that used it moved or for some reason or other doesn’t use it anymore, then it is really hard to just use it for something very different. So in the worst case scenario the microscope would not be used anymore. Of course one could just use the parts of the old one to build a new one for a different application, but then you also need a person willing to do that. The movement of OpenSPIM is just starting to arrive in the minds of biologists, so attempting to build your own is still somewhat rare. That said, the light-sheet microscopy and OpenSPIM community make it really easy to start into this adventure.”
About those data mountains…
The data output of a light-sheet microscope is several order of magnitudes higher than in conventional microscopy, says Gassler, making it necessary to invest in data storage and to explore ways to immediately reduce the data size. That can be done by omitting unnecessary data right after imaging or even during imaging sessions. And, she says, “to get the most out of light-sheet microscopy as a biologist, it is very valuable to team up with physicists and computer scientists.”

Tiago Pinheiro
PhD student in neuroscience and regenerative medicine
in the lab of Andras Simon in the
department of cell and molecular biology
Karolinska Institute
Stockholm, Sweden
{credit}Benny Coyac{/credit}
What he likes about light-sheet microscopy…
In his work with fixed, cleared salamander brains to study dopamine neuron regeneration, Tiago Pinheiro likes the speed with which images can be captured with light-sheet microscopy. Here is a video he made of stitched and processed images he generated on the ZEISS Z.1 microscope of glial protein fibers in the brain of a developing salamander. The brain had been cleared with Advanced CUBIC.
What is also beneficial about light-sheet microscopy, says Pinheiro, is being able to rotate the sample with just the right orientation. With confocal microscopy and 3D mounted samples that is a big hurdle. It takes many hands and much time to image a brain slice by slice to then find the paths of neurite fibers from slice to slice.
The advantage of OpenSPIM, in Pinheiro’s view, is that he can do experiments instead of waiting for the rather overbooked commercial light-sheet microscopes. If you know the OpenSPIM works for your specific application, he says, then a scientist could build several of these microscopes on a budget and speed up image acquisition for their experiment.

{credit}Vineeth Surendranath{/credit}
About being able to pack a microscope in a suitcase…
“It is great for education purposes,” says Pinheiro, who would love to have an OpenSPIM at Karolinska to show his colleagues and to make people aware of the potential of light sheet microscopy has and so they can see how it works. “As confocal microscopy made its way to every biology lab I am convinced light sheet microscopy will as well. A microscope in a suitcase is helping that happen.”
About the bigger scheme….
Research centers in biology and medicine have a growing need for staff with knowledge of physics and computing, says Pinheiro. A biologist can build an OpenSPIM after attending the EMBO course, as he has, but he or she will still need expertise at a home institution to trouble-shoot any issues such as assembly or software. More generally, he says not everyone will be able to take the course. But at the same time there is an urgent need to more quickly and extensively merge the fields of biology, computer science and physics. “I believe not doing that means falling behind in answering essential scientific questions in a better way,” he says.

Florian Vollrath
Physicist, programmer,
research associate in
the imaging facility at the
Max Planck Institute for Brain Research
Frankfurt, Germany
What he likes most about light-sheet microscopy…
Florian Vollrath helps scientists at the Max Planck Institute for Brain Research with their experiments and their data analysis. Vollrath and colleagues are ramping up to build a light-sheet microscope for the imaging facility. The model will have a different camera, stage and objectives than the basic OpenSPIM setup.
The institute mainly works with fixed samples where phototoxicity isn’t a problem but bleaching can be. What matter most about light sheet microscopy to him is its advanced measurement speed compared to confocal microscopes, he says. “Our dream is to image as fast as possible complete brains and being able to analyze their neuronal structure afterwards, without the need of slicing them in many pieces and imaging them one by one,” he says. Light sheet microscopes have a trade-off, their resolution is not as good as what can be achieved with confocal microscopes. “Our main question is now if it is still good enough.”
Being part of a community…
With an OpenSPIM community in place, it helps those with less or even no experience get on their way to working with light-sheet microscopy, says Vollrath. The open source software works, but it is not as advanced as the software in commercial systems, he says. It takes programming experience to adjust it if one wants to use components other than the ones on the OpenSPIM website.

Johannes Girstmair
PhD student
in the lab of Maximilian Telford
in the department of genetics, evolution
and environment
University College London{credit}Armin Märk{/credit}
About tapping into curiousity…
For biologists who are curious to get a start with OpenSPIM, Johannes Girstmair recommends taking one’s own samples to one of the around 70 OpenSPIM set-ups in labs around the world and finding someone who will “let you play around a little bit.”
About angles and speed …
“Speed does not always matter,” says Girstmair. It all depends on the question one is pursuing, he says. Speed matters with live imaging. For example he has looked at cellular behavior and cytoskeleton dynamics and tracked the nuclei of the developing embryos to create an early cell lineage. With a slow imaging system he can miss important information. He mainly uses one angle for time-lapse movies but time matters especially if someone is doing time-lapse live-imaging with multiple angles, “you don’t want to wait 2-3 min for each angle to be acquired simply because it would mean that with 5 angles you would need to wait almost 15 minutes per time-point,” he says. “A lot of development can happen in between.” And once the images from the previous angle are acquired, they risk not fitting well anymore with the acquired first angle. “That’s not good and might give you funny results once you fuse angles that are shifted in time quite a lot.”
He has also found that a faster, more smoothly running system can be better for the living embryos because a slow system may well delay the laser shutter, although he has not measured this, which means that the embryo might be exposed longer to the light-sheet, thereby increasing the chance of phototoxicity. If you can make a system faster and it does not cost much to do so, why not do it, he says.
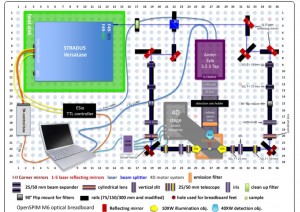
The configuration of the OpenSPIM model that Johannes Girstmair built.{credit}J. Girstmair{/credit}
About alignment…
Aligning the light sheet with the focal plane of the detection objective is tricky because the acquisition chamber has to be water tight. That limits the possibility of moving the detection objective that could otherwise be moved forward and backward to align the light-sheet well, says Girstmair. “We can cheat a bit by using the large corner mirrors to align the light-sheet to the focal plane,” he says. Information about how he assembled and aligned his system and videos of continuous imaging experiments are in his BMC Developmental Biology paper.
Also, he says, there are ways to nudge the detection objective a bit forward and backward in a way that the O-rings can tolerate and which are used to make the detection objective watertight. “People have a little wheel for this purpose, which doesn’t seem to be super hard to install if somebody insists on this,” he says.
About some questions that tempt him…
Girstmair studies evo-devo questions using, for example, the polyclad flatworm Maritigrella crozieri. These lophotrochozoans, or Spiralians as they are sometimes called, are interesting because so many phyla, including the flatworms, show a very similar developmental pattern early on, which is called spiral cleavage. This likely ancestral cleavage program allows scientists to compare the development of different phyla even though they have branched millions of years ago. Most flatworms don’t have a stereotypic spiral cleavage nor do they exhibit a free-swimming larval stage as are found in other lophotrochozoan phyla. M. crozieri has both the very stereotypic spiral cleavage pattern and a free-swimming planktotrophic larval stage, says Girstmair, making these embryos a good starting point for comparative studies.
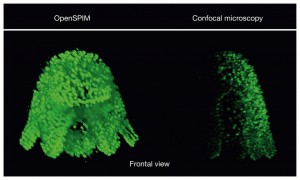
The polyclad flatworm Maritigrella crozieri imaged with different techniques{credit}J. Girstmair{/credit}
About needing a ‘Pavel’…
Pavel Tomancak, one of the co-founders of OpenSPIM, is a co-author on Girstmair’s paper about building OpenSPIM and using it to study Maritigrella. Tomancak’s presence might make his project look a little less like a do-it-yourself one. “Of course not everybody can have a ‘Pavel’ close by,” as he did, says Girstmair. But for starters they can travel to a lab with an OpenSPIM set-up and work there with their own samples.
“As for the assembly in London I really put everything together myself and more importantly hardware-configured the microscope myself,” says Girstmair. Several people offered plenty of advice, which is why, he says, they also deserve to be on the paper. They include Tomancak, former Tomancak lab member Peter Pitrone now a light-sheet microscopy consultant and Mette Handberg-Thorsager, a developmental biologist also in Dresden with whom Girstmair tested microinjection techniques.
For the OpenSPIM setup, Girstmair and his colleagues used some parts that differ from the basic set-up such as a multi- laser system, controller boxes and other components, which also meant there was “a lot more to learn and sometimes even get frustrated about,” he says.
This OpenSPIM image comes from a fixed embryo and imaged with multiple views. The nuclei of each cell are stained with the nucleic acid stain SytoxGreen. The first angle will also be the orientation of the 3D reconstructed embryo when all the different views are combined into a single image file using software called Fiji.
“I think the images made with the OpenSPIM are not particularly better than the confocal images,” says Girsstmair. The confocal images are crisper and have better resolution. But they can’t contain all the information contained in an image captured from multiple angles. Imaging at multiple angles is very difficult with a conventional confocal microscope due to the different ways specimens are mounted, he says.
Fixed specimens imaged with OpenSPIM are usually embedded in agarose and therefore keep their natural shape. “With the confocal I would try to squeeze a stained Mueller’s larva as much as possible in order to get the most out of the staining from a single view and thereby I also loose the specimen’s natural shape,” he says. When it comes to capturing the development of Maritigrella embryos, OpenSPIM is much better: it is faster and the embryos are exposed to much less light. Another advantage: the freely available software tools for 3D reconstructions.
OpenSPIM is a crowd-sourced movement propelled by the crowd, among them, these people:

{credit}Vineeth Surendranath{/credit}
Peter Pitrone (top), is first author on the paper presenting OpenSPIM, Pavel Tomancak (third from the top), a researcher at Max Planck Institute (MPI) for Molecular Cell Biology and Genetics in Dresden co-developed OpenSPIM. The others in this photo are PhD students who took a course on OpenSPIM and who put together the OpenSPIM web site.