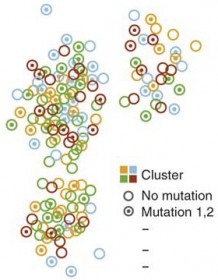
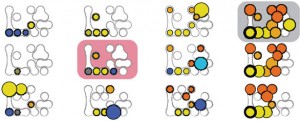
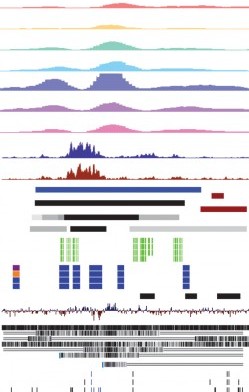
Box plots are excellent for visualizing important core statistics of sample data. We hope that a new online plotting tool BoxPlotR will help encourage their wider use in basic biological research.
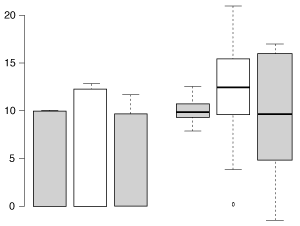
The same three samples plotted by bar chart with s.e.m. error bars (left) and Tukey-style box plot (right). The box plot more clearly represents the underlying data.
Box plots are heavily used in biomedical research in which statisticians have historically had considerable input into study design and analysis. But although similar types and quantities of sample data also appear in basic research (such as that published in Nature journals) box plots are much less common than bar charts in these manuscripts. Last year in Nature Methods for example, ~80% of sampled data was plotted using bar charts.
Discussions we had with the community suggested that an impediment to using box plots instead of bar charts to graph sample data was due to limited support for box plots in plotting programs commonly used by researchers. It also became apparent that some software that did support the box plot was deficient in communicating to users what the different elements of the plot represented. As a result, strangely labeled box plots were showing up in published papers. At NPG we thought it would be useful to provide authors with a simple online tool they could use to generate basic box plots of their data for publication.
The origin of BoxPlotR
At the VizBi 2013 conference in Cambridge Massachusetts I mentioned NPG’s desire for such a tool at a breakout session chaired by Martin Krzywinski in which the participants, including a young researcher named Jan Wildenhain, discussed what the community needed to create better figures. I also happened to mention our interest in this to Michaela Spitzer while visiting her poster from the Juri Rappsilber and Mike Tyers labs showing how the R-package ‘shiny’ by RStudio can be used to easily convert R code (a popular scripting language for statistics) into a visual application for exploring data.
Later at the conference Jan approached me and said he was intrigued by our desire for someone to design a webtool to create box plots and that he was interested in working on such a project. I happily told him to get in touch with me after the conference so we could discuss it further.
Three weeks after the conference concluded I still hadn’t heard from Jan and was beginning to worry that he had decided not to pursue this. Then… a few days later, I received an email from Jan. Much to my surprise he provided a link to a highly functional tool that he and Michaela, through their own initiative, had gone ahead and created using shiny and R. What followed was a productive and rewarding period of discussion and development during which time Michaela incorporated additional functionality and made selected design changes. The tool appeared so well designed and functional that I encouraged them to submit it to Nature Methods for publication as a Correspondence. After incorporating additional functionality and changes based on comments brought up during peer review BoxplotR was ready for publication.
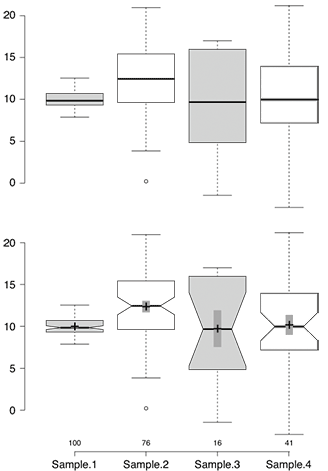
Sample BoxPlotR plots. Top: Simple Tukey-style box plot. Bottom: Tukey-style box plot with notches, means (crosses), 83% confidence intervals (gray bars; representative of p=0.05 significance) and n values.
To accompany the publication and launch of BoxPlotR we thought it would be useful to provide some information and practical advice about box plots to our readers. Nils Gehlenberg, a former author of several Points of View articles with Bang Wong, agreed to resurrect that popular column for our February issue with an article on bar charts and box plots. Similarly, Martin Krzywinski and Naomi Altman agreed to delay our planned Points of Significance article on the two-sample and paired t-test and instead devote an article to box plots.
Seeing how the community responded to our interest in creating an online box plot tool and then working with them on this project has been a great experience. This never would have been possible without the initiative and talent of Jan and Michaela or the support they received from their PIs Mike and Juri. We hope both our authors and others find BoxPlotR useful and we encourage feedback. General comments can be made here on our blog or by emailing the journal. For specific bug reports and feature requests please see the contact information at https://boxplot.tyerslab.com.

 Elements of a figure
Elements of a figure