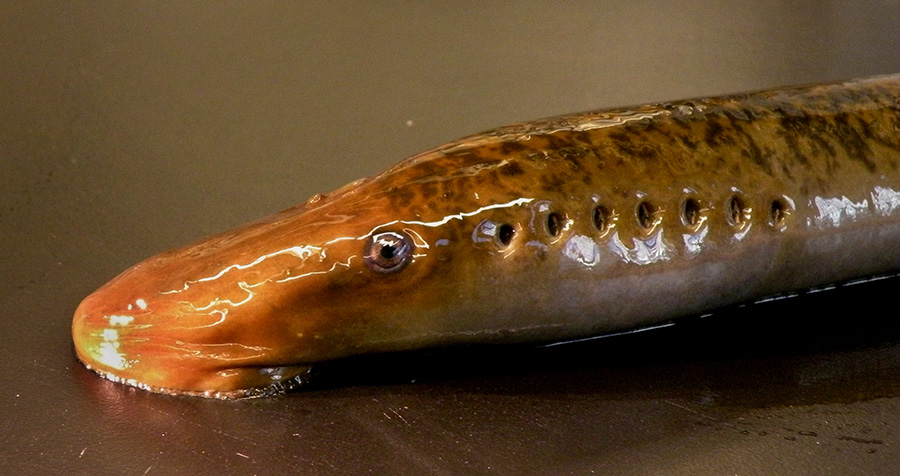
The sea lamprey (Petromyzon marinus) is an important model in evolutionary biology. It was discovered in 2009 (https://www.pnas.org/content/106/27/11212.long) that the genome of the sea lamprey undergoes extensive programmed genome rearrangement during development, where ~0.5 Gb (around 20%) of DNA is eliminated from the genome. The somatic tissues contain smaller genomes and only the germ cells retain the full complement of genetic material. The genome of the sea lamprey had been sequenced previously from the blood and liver, so only the somatic genome has been thoroughly characterized (https://www.nature.com/articles/ng.2568).
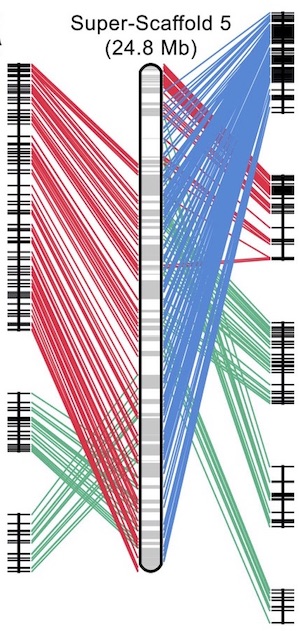
In a paper published this week in Nature Genetics, Jeramiah Smith and colleagues report the germline genome sequence of the sea lamprey. Using a combination of shot-gun and long-read sequencing integrated with scaffolding data and a meiotic map, the authors assembled a high-quality genome with near-chromosome level of contiguity. This allowed them to identify hundreds of genes that were systematically eliminated from the genome during development. Comparative analysis showed that mouse homologues of these genes are often marked by repressive complexes, indicating parallel strategies for programmed development.
We spoke with lead author Jeramiah Smith from the University of Kentucky to get some background on this research:
- What inspired you to sequence the germline of the sea lamprey?
I have worked with lamprey for years. I originally got involved with lamprey because it holds a special place in the vertebrate tree of life that shed light on the common ancestor of all vertebrates. That was the motivation for the first lamprey genome project, which sequenced DNA from blood and liver cells. Once we started working with lamprey we found out that the genome was much more complex than we ever anticipated. This included the fact that the genome changes its sequence content in a reproducible manner over the course of its normal development: something we call programmed genome rearrangement. The amount of DNA that is eliminated from sea lamprey is more than is present in some entire fish genomes, roughly half a billion bases. For me, this finding was the major inspiration behind sequencing the germline genome.
- What do you think were the most surprising or interesting findings to come out of the sequencing?
There were quite a few, but the strong overlap between programmed genome rearrangement and Polycomb-mediated silencing was near the top. The other was the rather strong evidence suggesting the some chromosomes, including chromosomes carrying the HOX genes, appear to have duplicated rather recently and seemingly independently from the rest of the genome. It’s a really strange genome.
- Can you comment on programmed genetic elimination as a developmental strategy versus Polycomb-mediated silencing?
Polycomb-mediated silencing arose deep in our evolutionary history, and is even present in unicellular organisms. We know that lamprey possesses human homologs of all Polycomb genes, but also uses programmed elimination. The difference between programmed elimination and other mechanisms of gene silencing is that programmed elimination is essentially irreversible, given that the DNA is physically removed. This means that the genes can never be expressed after an embryonic cell lineage has undergone elimination. Other silencing mechanisms are generally reversible, meaning that gene expression can be reactivated. In some cases reactivation is important. For example, in the context of development and regeneration. But in other cases activation of genes in the wrong tissue can case diseases, such as cancer. Lamprey seems to know which genes should never be reactivated outside of the germline.
- What is the most challenging part about working with sea lamprey?
The Genome! Aside from undergoing complex changes during development it also contains a large amount of repetitive DNA and a lot of sequence polymorphism. These features present substantial challenges for assembly and downstream analyses, but we’ve found that they can also be useful tools. We’ve used the abundance of sequence polymorphisms as a tool for mapping genes in lamprey and we now think that some classes of repeats are going to be critical for our future work aimed at figuring out how eliminated DNA is identified and packaged in the early embryos. Lampreys also only breed once a year and take from 5 to maybe 20 years to mature, this makes some experiments impossible, but lamprey researchers are very creative and the community has figured out how to get a lot done in this system.
- What organisms would you like to see sequenced in the future to help resolve the evolutionary relationships of vertebrates?
There are so many! Hagfish are going to be critical. They are another deep lineage that provides important perspective on vertebrate evolution and also happen to undergo programmed DNA elimination. There are also two other deep lamprey lineages that I also think will be important. Those species live in the southern hemisphere and diverged from sea lamprey around 300 million years ago, as opposed to the roughly 600 million year divergence between lampreys and other vertebrates. A lot of evolution can happen over 600 million years and these species should help bridge that gap. Salamanders and other amphibians are also going to fill important gaps and teach us a lot about the way vertebrate genomes evolve and function. It also seems certain that new sequencing technologies are also going to give us better genomes for other important species that have already been sequenced (e.g. amphioxus, sharks and shark relatives, and even sea lamprey). Finally, I think the zebrafinch germline genome will also be really interesting. They seem to have recently evolved something similar to lamprey’s programmed eliminations, and have a chromosome that’s unique to their germline. I’d really like to know what’s on that chromosome.