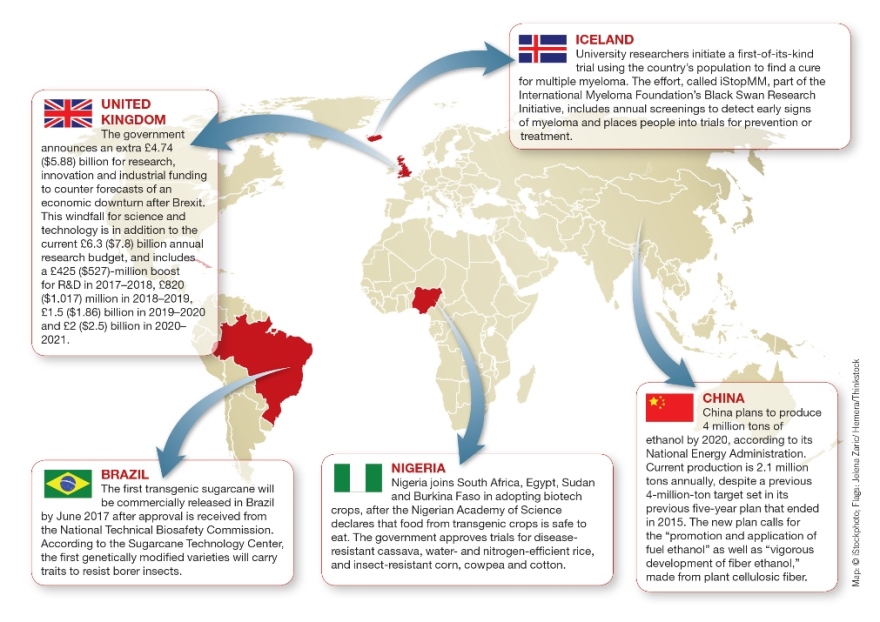
Chinese ethanol production levels, an Icelandic multiple myeloma initiative, Brazilian transgenic sugarcane, and more in January’s Monthly Map. (click to enlarge)
Author Archives: Bioentrepreneur
Medicago’s proteomic profile
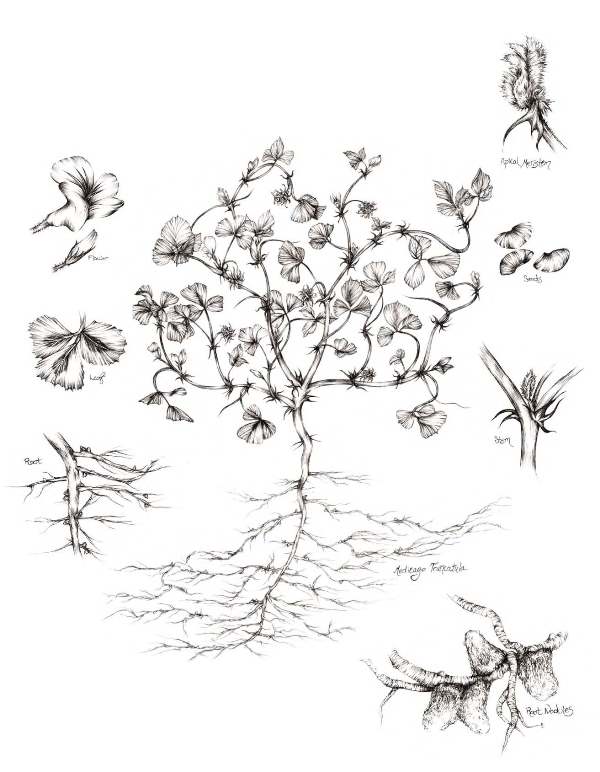
Legumes have the unusual ability to make their own fertilizer. They do this by associating with nitrogen-fixing bacteria, which take nitrogen from the air and convert it to ammonia that can be used by the plant to make proteins, nucleic acids and other essential molecules. Understanding the biology underlying the association between legumes and their symbionts could enable these concepts to be adopted in other types of crops as well. In a paper recently published in Nature Biotechnology, Joshua Coon and colleagues report the proteomic profile of the organs of a model legume, Medicago truncatula, and its collaborative partner, Sinorhizobium meliloti. Analyzing the proteomic atlas they generated together with transcriptome and genomic data, the researchers show how this resource can be used to discover new biological insights surrounding nitrogen fixation that may one day be useful for future crop engineering efforts.
Monthly Map
Mitochondrial protein functions
Mitochondria are energy-generating organelles that carry a portion of their own DNA. Mutations in mitochondrial proteins are associated with metabolic disorders, neurodegenerative diseases and a variety of malignancies. Joshua Coon, David Pagliarini and their colleagues generated proteomic, lipidomic and metabolomic data on 174 strains of yeast, each carrying a mutation in a different protein implicated in mitochondrial function. In their paper in the November issue of Nature Biotechnology, they show how connections among these different data sets can be exploited to reveal new insights into the biosynthesis of coenzyme Q, an essential lipid required for oxidative phosphorylation. Their paper can be found here.
Bio-leaders of Tomorrow
 I first came across the GapSummit in November 2015. The meeting was organized by Global Biotech Revolution and was to be held April 2016 at Cambridge, UK. Looking at the agenda, I noted the Voices of Tomorrow competition, which aimed at getting all the participants to form groups and come up with possible solutions for the ‘gaps’ in the biotech sector, found in research and innovation, funding, future health, future resources, people, bioethics, and public perception & education.
I first came across the GapSummit in November 2015. The meeting was organized by Global Biotech Revolution and was to be held April 2016 at Cambridge, UK. Looking at the agenda, I noted the Voices of Tomorrow competition, which aimed at getting all the participants to form groups and come up with possible solutions for the ‘gaps’ in the biotech sector, found in research and innovation, funding, future health, future resources, people, bioethics, and public perception & education.
I applied to attend as a leader of tomorrow. This required me to write an essay explaining why I wanted to attend and in December, I was informed that I have been selected to attend the Gapsummit 2016. I was elated!
The “leader of tomorrow” title I think is apt, since the science students of today will be future leaders. Millions of students and postdocs like me are future leaders in the biotech sector. We are the ones who are going to make life liveable in this fast-paced world!
In April, I attended the GapSummit 2016, and it was a phenomenal experience. According to their website, “GapSummit is the world’s first inter-generational and inter-cultural leadership summit in biotech.” It aims “to connect the biotech think-tanks, industrial leaders and research pioneers who are going to become the young bio-leaders of tomorrow and initiate a discussion about the gaps in the sector.”
This was definitely what I experienced when I was at the meeting. I found the experience enriching and felt the need to do a similar event in Singapore, the first of its kind here. I wanted to create a conducive atmosphere for groups of focused and motivated bio-leaders who will help advance the biotech and healthcare ecosystem in Singapore.
After several discussions with Global Biotech Revolution, they decided to organize the Singapore Leaders of Tomorrow (SLoT) Forum (thanks to Kelvin Chan for facilitating this!), to be held on October 19th 2016, with Biotechin.Asia (Disclaimer: I am the co-founder of Biotechin.Asia) as a key partner. I then started forming a team in Singapore; I roped in Laxmi Iyer, my partner at Biotechin.Asia, and other colleagues to join in as part of the executive team. A*STAR, where I am a research fellow, has been supportive of this effort, and the forum will be held at their premises.
The summit should help Singapore form more science competitions, and could provide pointers on teaching the next generation of bioscientists.
Monthly Map
Analyzing Global Biotech Investing Over Time
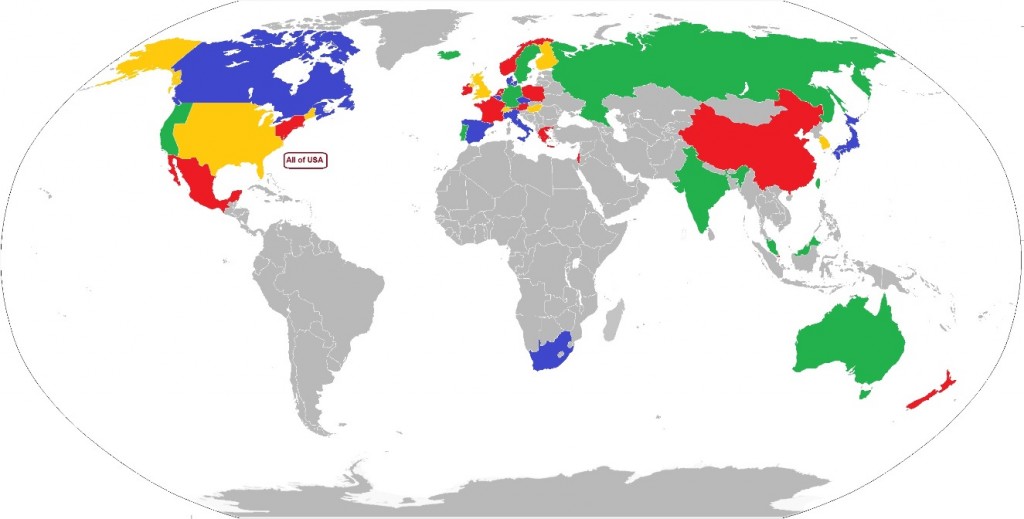 Where are venture funds being deployed in biotech? Is this changing? As part of a programme to find out how VC interacts with biotech, I have created a database of venture investment in 33 countries and three regions of the USA, over the time periods before, during and after the financial crisis of 2008-11, and built this into an interactive map of the biotech VC world. The map is clickable, with data for each country linked to that country on the map. The link to the map is here.
Where are venture funds being deployed in biotech? Is this changing? As part of a programme to find out how VC interacts with biotech, I have created a database of venture investment in 33 countries and three regions of the USA, over the time periods before, during and after the financial crisis of 2008-11, and built this into an interactive map of the biotech VC world. The map is clickable, with data for each country linked to that country on the map. The link to the map is here.
A quick summary shows:
- USA dominates investment in biotech.
- East and West Coast investors have different behaviour and different investment priorities than others.
- UK, Israel and non-Coastal US investors are similar
- Looking to the Coastal US for examples of ‘how to do it’ guides for policy may not work.
- Companies with bold ambitions should look to where investors match their vision, not to local sources.
Globally, the USA continues to dominate investment in biotechnology – around 2/3 of all investments by value and number are from investors in the USA and into US companies.
Because of the US dominance, many European governments and a not insignificant number of entrepreneurs look to the USA for leadership, inspiration and, of course, funds. But is the USA a good model for other countries?
To MBA or not to MBA: is that the question?
“What kind of CEO do you want to be?” That question still rings in my head.
It came after a meeting with a CEO that I’d attended with my dad, one in a string of meetings to match small biotech and health tech companies with potential investors.
My dad, an investor, was letting me flex some muscle. I had hopped straight from earning a PhD in biomedical engineering to consulting at McKinsey – and now, I knew just the questions to ask: if it was a cell therapy or biologic, I’d hammer them on manufacturing or the regulatory process. If it were a device, I’d ask about IP or competition.
Over time, I noticed a pattern in the answers. If the CEO were, say, an MD, he or she would demonstrate less fluency with the financials. And if it were a lawyer at the helm, I’d note the micro-stuttering of a person for whom technology was a “second language” – in other words, someone who’d been taught (rather than having developed) the technology’s mechanism of action.
What I was intuiting from those meetings, it turns out, was the weak spot common to any technologically-based venture: the gap between technology and management, between science and business. This is a language barrier as formidable as any other. Like the gaps between other spoken languages, it blocks understanding within biotech companies, impedes their progress, and is difficult to overcome. Indeed, the deeper a company’s chasm, the more reluctant my dad was to invest.
Guidelines for algorithms and software at Nature Biotechnology
Nature Biotechnology outlined its recommendations for material sharing and reporting for algorithms and software in a April 2015 Editorial. On this page, the editors provide more detailed information for authors submitting manuscripts containing unpublished algorithms and software.
Client-side Software
This is software that is installed and used on a personal computer and not intended to be accessed remotely as a web service. It can be entirely stand-alone on a commonly available operating system (Windows, Mac OS X, or *nix) or can require the user to have a popular software platform installed (MATLAB or LabVIEW). In all cases, but particularly when using MATLAB or LabVIEW, all platform versions and software dependencies must be detailed in the supplied documentation.
At Submission
- If the custom algorithm/software is central to the method and has not been reported previously in a published research paper it must be supplied by the authors in a usable form including one or more of the following.
- Source code
- Complete pseudocode
- Full mathematical description of the algorithm
- Compiled standalone software