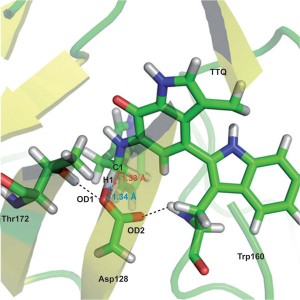
At Nature Chemistry we all love Density Functional Theory and we all love polls – so what could be better than a poll on DFT? Marcel Swart, Matthias Bickelhaupt, and Miquel Duran have, for the last five years, been running a poll to find out which functionals the computational chemistry community like or dislike. The results of the the 2014 poll are now out and we have a guest post from the three of them to explain a little more about it.
I’m sure we’ll be seeing this as a question on Family Fortunes/Family Feud soon: “We asked 100 people to name …. a popular density functional”
Gavin
(Senior Editor, Nature Chemistry)
*******************************
Since 2010 we have been organizing an annual online popularity poll for density functionals where we probe the preferences of the computational chemistry community for their preferred Density Functional Theory (DFT) functionals. It all started with a presentation by Matthias Bickelhaupt (Feb. 2009) that showed the values of various chemical properties calculated using quite a number of different density functionals. Miquel Duran suggested, essentially, ‘averaging’ a number of these values but with appropriate weightings applied that reflected how ‘good’ the employed functionals were. Thus obtaining a ‘consensus’ density functional result. This could then act as a measure of how well the computational chemistry community is doing if compared to state-of-the-art reference data.
In order to get the weightings needed for this procedure, we have held annual online polls where people could indicate their preferences for a number of density functionals. The polls were announced on the Computational Chemistry List (CCL; a mailing list where people can ask for advice about any aspect of computational chemistry), on Twitter, Facebook, blogs, and so on, in order to get the maximum number of participants. The aims of this poll were: (i) to probe the ‘preference of the community’, that is, setting up a ranking of preferred DFT methods; and (ii) provide a compilation of the ‘de facto quality’ that this implies for the ‘average DFT computation’.
The DFT poll has led to, what some might deem to be, a polarization of the field, with some people clearly in favor (e.g. Steven Bachrach, Gernot Frenking, John Perdew, Henry Rzepa), and others clearly against. Especially this year there has been a vivid debate on the CCL mailing list in the days after the poll was announced (CCL June 1st entries, CCL June 2nd entries, CCL June 3rd entries). This may, or may not, be related to the fact that in 2013 we had to disqualify one functional that had fallen victim to a blatant attempt to bias the outcome of the poll. This has not happened again this year (because we switched to another survey provider).
In order to stress the motivation for holding the poll, we as organizers felt we needed to add a statement as well, since we “are simply monitoring what happens in the field of DFT and comment on how the choice of the community differs from (or agrees with) reliable reference data. In that way, we do exactly what should be done, namely ‘drive science through evidence and logic’ or maybe even ‘drive science back to evidence and logic’ (because, against all basic principles of science, the community often just follows blindly a fashion)”. And as one message nicely described: “Yes, it is not scientifically sound, epistemologically correct, platonically unsullied. But at least it is fun. We should appreciate fun in chemistry”.
The nice part of the popularity poll is that it opens the field to newbies, who through the poll have a better understanding of which are the most popular functionals within the field — which can serve as a good starting point for those looking for the best functionals for their given problems. For instance, the rise of the wB97X-D functional is nicely reflected in this year’s results (moving upwards to 4th place after PBE, PBE0 and B3LYP), as was the fact that the winner of the first editions (PBE0) was largely unknown to many people. The years after the first edition (in which it ‘won’), the number of PBE0 citations has increased considerably, by at least 60% (see Figure 1) for the three PBE0 papers (there is the original paper by Perdew and co-workers where they describe the rationale for using 25% Hartree-Fock exchange; and there are two separate papers that introduce the functional as being PBE0, by Ernzerhof and Scuseria, or by Adamo and Barone).
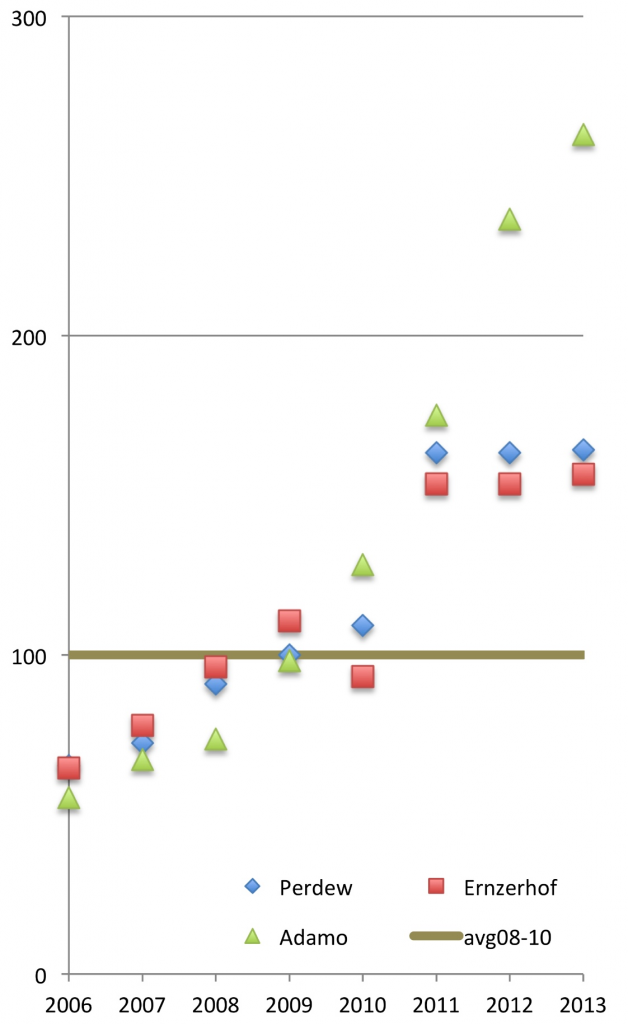
Figure 1. Normalized number of citations for PBE0 papers, before (2006-2010) and after (2011-2013) the first news-item of the DFT poll (100 = average number of citations for 2008-2010 for each of the three papers separately).{credit}Courtesy of Marcel Swart{/credit}
Given that this is the fifth edition, we asked several researchers in the field (in Sept. 2014) whether they were in favor or against the DFT poll.
Steven Bachrach: “Please feel free to quote me from the Wiley Interdisciplinary Reviews article [“It would be nice if we could somehow again reach some consensus regarding a uniform standard computational method that experts and non-experts could rely upon for most situations. A challenge I make here to the computational community is to try to reach an accord on establishing a standard methodology. Perhaps a conference could be called where leaders propose their best methods and after discussion, a vote yields a recommendation for the greater user community”] and from CCL [“I also think the poll has value in discerning trends, especially new functionals to appear on the list and ones that have fallen down or off”].
John Perdew: “The DFT popularity poll is somewhat like citation analysis: It measures (but in a different way) how well a functional has been received by a set of readers and users. There are many reasons why some functionals are received better than others: accuracy, reliability, wide applicability, computational efficiency, well-founded construction, availability in standard codes, reputation of the functional and its authors, historical priority, novelty, and even hype. The poll has to be seen as measuring all these things, and perhaps more. To the extent that the polled scientists use rational criteria, the results of the poll can point other scientists toward good or interesting functionals”.
Henry Rzepa: “I still think the context of any vote cast is absolutely crucial. Perhaps what the community needs to develop is a public set of conformance test sets of molecules, one for each type of property?”
Andreas Savin: “I must shamefully confess that I do not know about the DFT popularity poll.” (after having received more information): “I will not participate, as this poll is intended for people who apply DFT, and I do little in this direction, but I find it interesting. I am amused to see that B3LYP is not as popular as generally believed, and LDA has such a high rank. How does it compare to the number of citations?”
Gustavo Scuseria: “I am not in favor or against the poll. It is interesting though that we need a contest to determine what is popular and useful. A cacophony of functionals have mushroomed in recent years, and I am very much afraid that uncontrolled approximations and rampant empiricism have taken over DFT”.
An interesting addition was brought forward in the discussions this year by Henry Rzepa (June 1, 2014), who suggested that, in the future, the poll should be extended to enable participants to explain why they like or dislike a given functional. This is a new interesting feature that will be added to next year’s edition: for each functional the participants can indicate for a number of properties whether they love using the functional, or rather dislike it. Rzepa proposed a number of properties (reaction barriers, normal mode analysis, NMR shieldings, etc.), which will be fine-tuned before the polling season opens again on June 1 2015!