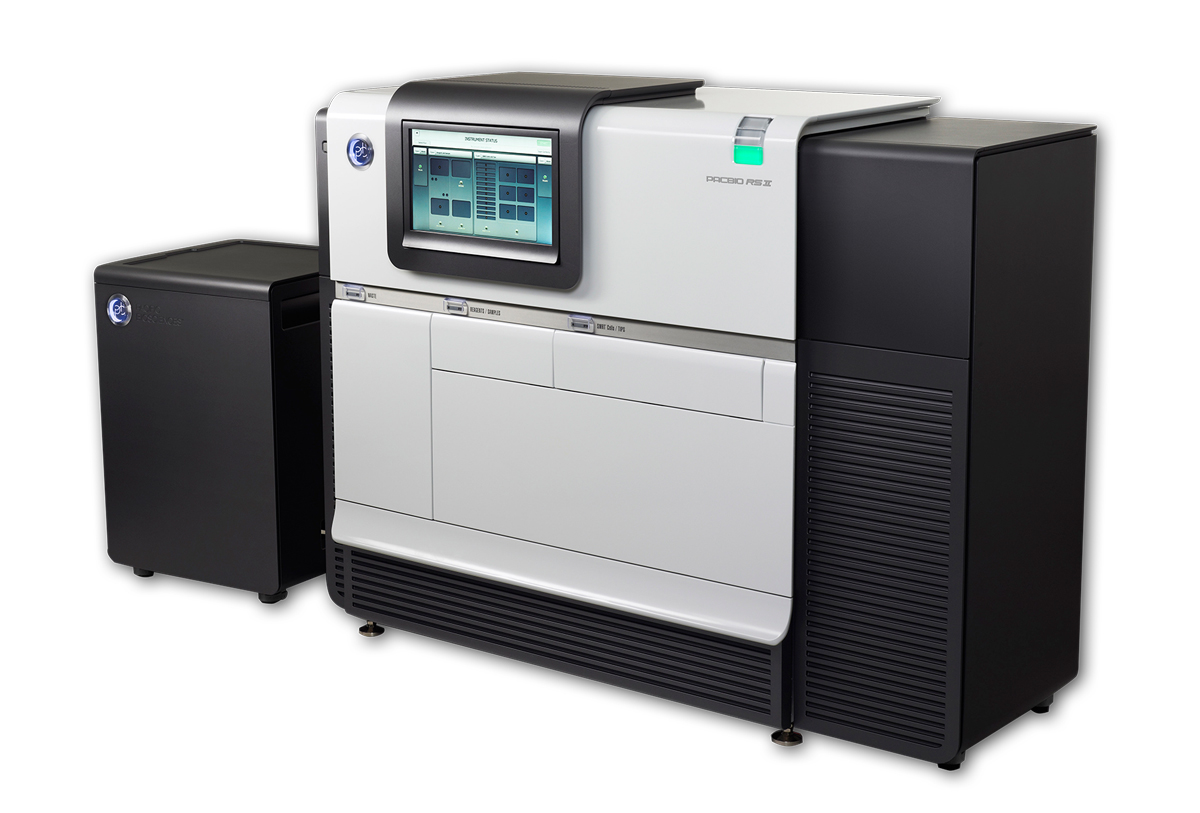
Sometimes, a DNA sequencer is more than it seems. In this month’s Technology Feature, I talk to the researchers who have figured out ways to squeeze new life from an outdated DNA sequencer, the Illumina GAIIx. That’s a popular choice for sequencer-hackers, but not the only one. Stanford structural biologist Joseph Puglisi uses a PacBio RSII from Pacific Biosciences to plumb the biochemistry of protein translation.
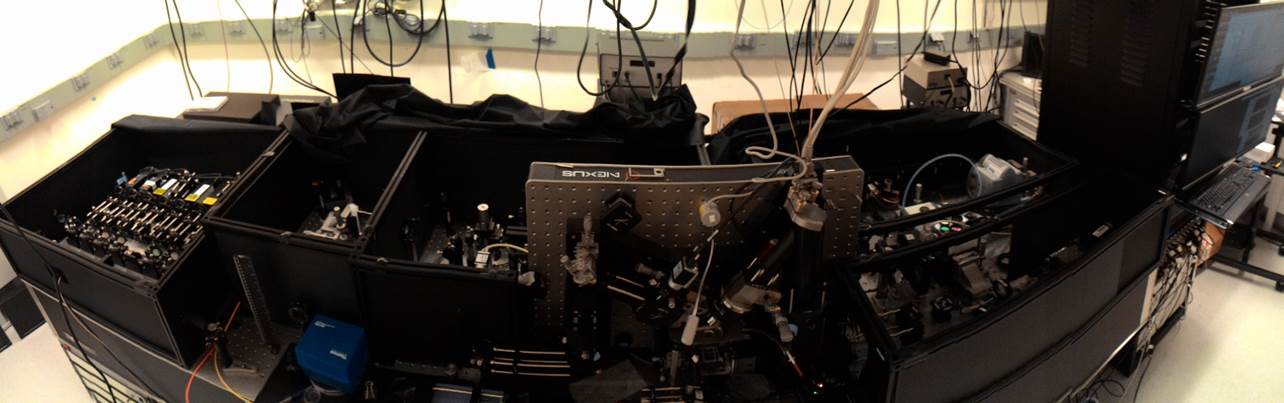
The RSII was designed as a single-molecule DNA sequencer, in which powerful cameras capture the flashes of light that result when a DNA polymerase molecule tethered to the base of a microscopic well inserts a fluorescently labeled base into newly synthesized DNA. But according to Jonas Korlach, the company’s chief scientific officer, that’s just one of its applications. “Yes, it’s a sequencer, but at the same time it’s also the world’s most powerful single-molecule microscope.”
All that’s required to make that microscope record something other than DNA synthesis, fundamentally, is for researchers to replace the tethered DNA polymerase with another enzyme, and to add the appropriate fluorescent reagents. To alter the running conditions, researchers also need PacBio to ‘open’ its system software to afford them greater control — for instance, to adjust experimental temperature, imaging conditions, and fluid addition. According to Korlach, just four instruments worldwide have been tweaked in this way. (As with the Illumina hardware discussed in the Technology Feature, such hacks only work on PacBio’s older RSII; the newer Sequel is not hackable, Korlach says.)
The company offers these researchers what support it can, but because they are pursuing home-brew applications, Korlach says, researchers who run into technical issues must solve them in-house. “They are mostly on their own.”
Researchers have used these modified systems to address the biophysics of cell-cell interaction, transcription, splicing, and in Puglisi’s case, translation. Puglisi’s is a structural biology lab, and structural methods tend to provide static pictures. But biology is dynamic. So, his team typically pairs the methods up. “We always like to couple structural investigations with some way to animate the structure and bring it to life,” Puglisi says. Since 2014, the lab has published some 25 studies using the RSII to study the ribosome.
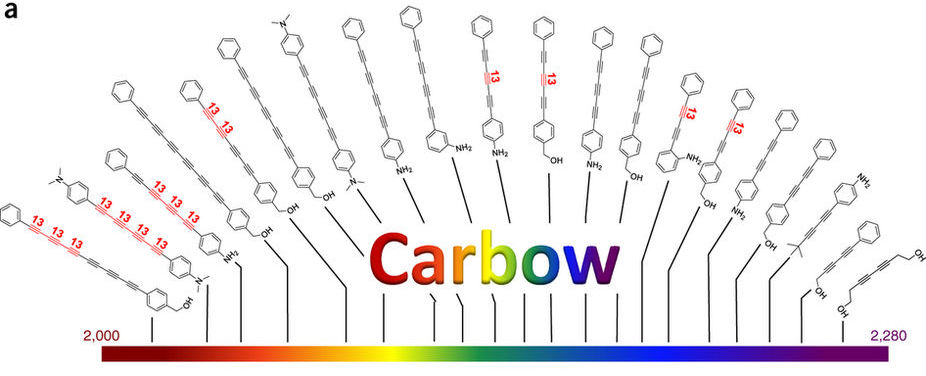
In one recent study, for instance, Puglisi’s team studied the impact of modifying one particular carbon atom in the backbone of RNA. That modification, they found, causes the ribosome to pause, possibly in order to allow ancillary biological processes, such as protein folding or protein processing, to occur.
“The biology of the system really still needs to be worked out, but the dynamic behavior and structural signatures that we saw were so striking that … there has to be some neat biology here,” Puglisi says.
Korlach, who worked with Puglisi on some of his earliest efforts on the RSII, says the team, with Puglisi’s postdoc Sotaro Uemura (now at the University of Tokyo) worked out these methods on nights and weekends, when the laboratory was otherwise unoccupied. And he recalls the excitement of getting the system to work that first time.
“It was pretty thrilling when we saw the first traces of real-time dynamics of ribosome translation,” he says. “That was the first time any human had ever seen a ribosome make a protein in real time on a single-molecule level, with codon resolution. Those are the types of milestones that as a method developer you live for.”
Jeffrey M. Perkel is Technology Editor, Nature
Suggested posts
Lattice light-sheet microscopy gets an AO upgrade