This month we launch a new column ‘Points of Significance’ devoted to statistics, a topic of profound importance for biological research, but one that often doesn’t receive the attention it deserves.
For the past three years Nature Methods has been publishing the Points of View column, one page a month dedicated to practical advice for researchers on how to create accessible and accurate visualizations of their data. The response to the column articles has been fantastic and most recently we organized them by topic here on our blog.
Unfortunately, a truth about data visualization is that no matter how good the visualization, if the experiment wasn’t appropriately designed and the data wasn’t analyzed correctly, the resulting visual depiction of the data will be inherently flawed. Nature Methods and the other Nature journals recently made changes to improve data and methods reporting as part of a reproducibility initiative. We feel this is an important first step in improving experimental reproducibility and repeatability, but unfortunately by the time work is submitted for publication it can be difficult to correct shortcomings in experiemntal design and analysis.
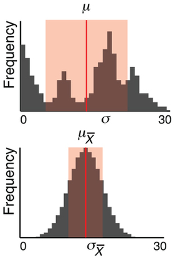
A population distribution and a distribution of sample means.
In our September issue readers will find a new column, Points of Significance, that we hope will be as useful as the column that preceded it, perhaps more so. Martin Krzywinski, who has been writing the visualization column, is now joined by Naomi Altman, Professor of Statistics at The Pennsylvania State University. Among other things, Naomi will be responsible for ensuring that the information and advice we provide about statistics in every Points of Significance article is accurate.
The column has been expanded from one to two pages and will often have an Excel spreadsheet associated with it. This expansion will help us better communicate information that is less well served by display items. However, as illustrated by the figures in the first article of the column and the accompanying spreadsheet, visual displays will continue to play a vital role due to their strength in providing easily interpretable examples that can often be more readily grasped than mathematical or narrative descriptions.
We will strive to present the material so that each article in the column builds on prior ones. In this spirit the first article discusses populations and sampling, a foundation for nearly all topics to follow. The accompanying spreadsheet allows readers to play around with sampling and see for themselves how often values obtained from samples deviate substantially from the real population. It can be disconcerting to see just how often ‘bad luck’ can give a ‘wrong’ result in one set of measurements while in another set of measurements the ‘right’ result is obtained but statistical measures would suggest that the former is more likely to be ‘correct’ than the latter. This excellently highlights how statistics is unable to tell you if you are right. But this doesn’t suggest statistics has limited value. Instead, readers of scientific articles reporting statistical results need a healthy grasp of the limitations of statistical analysis and users of statistics can always learn ways to improve the power of their analysis.
The “aura of exactitude” that often surrounds statistics is one of the main notions that the Points of Significance column will attempt to dispel, while providing useful pointers on using and evaluating statistical measures. We expect that readers will find the upcoming October Points of Significance article on error bars and confidence intervals with its practical tips on interpreting these graphical elements to be particularly useful almost every time they read a manuscript containing these popular visual representations of uncertainty.
We hope readers enjoy Points of Significance. It is appropriate that the column is debuting during the International Year of Statistics. To allow readership by a wider audience each article will be free to access for a period of one month after it is published.
Update: All Points of Significance articles are now free access and have been collected together on a dedicated page in the nature.com “Statistics for biologists” resource.
For more on statistics, and particularly statistics training, don’t miss this September’s Editorial.
. . . . . . . .
Update: Below is a continuously updated list of the Points of Significance articles.
Importance of being uncertain – September 2013
How samples are used to estimate population statistics and what this means in terms of uncertainty.
Error Bars – October 2013
The use of error bars to represent uncertainty and advice on how to interpret them.
Significance, P values and t-tests – November 2013
Introduction to the concept of statistical significance and the one-sample t-test.
Power and sample size – December 2013
Using statistical power to optimize study design and sample numbers.
Visualizing samples with box plots – February 2014
Introduction to box plots and their use to illustrate the spread and differences of samples.
Comparing samples—part I – March 2014
How to use the two-sample t-test to compare either uncorrelated or correlated samples.
Comparing samples—part II – April 2014
Adjustment and reinterpretation of P values when large numbers of tests are performed.
Nonparametric tests – May 2014
Use of nonparametric tests to robustly compare skewed or ranked data.
Designing comparative experiments – June 2014
The first of a series of columns that tackle experimental design shows how a paired design achieves sensitivity and specificity requirements despite biological and technical variability.
Analysis of variance and blocking – July 2014
Introduction to ANOVA and the importance of blocking in good experimental design to mitigate experimental error and the impact of factors not under study.
Replication – September 2014
Technical replication reveals technical variation while biological replication is required for biological inference.
Nested designs – October 2014
Use the relative noise contribution of each layer in nested experimental designs to optimally allocate experimental resources using ANOVA.
Two-factor designs – December 2014
It is common in biological systems for multiple experimental factors to produce interacting effects on a system. A study design that allows these interactions can increase sensitivity.
Sources of variation – January 2015
To generalize experimental conclusions to a population, it is critical to sample its variation while using experimental control, randomization, blocking and replication to collect replicable and meaningful results.
Split plot design – March 2015
When some experimental factors are harder to vary than others, a split plot design can be efficient for exploring the main (average) effects and interactions of the factors.
Bayes’ theorem – April 2015
Use Bayes’ theorem to combine prior knowledge with observations of a system and make predictions about it.
Bayesian statistics – May 2015
Unlike classical frequentist statistics, Bayesian statistics allows direct inference of the probability that a model is correct and it provides the ability to update this probability as new data is collected.
Sampling distributions and the bootstrap – June 2015
Use the bootstrap method to simulate new samples and assess the precision and bias of sample estimates.
Bayesian networks – September 2015
Model interactions between causes and effects in large networks of causal influences using Bayesian networks, which combine network analysis with Bayesian statistics.
Association, correlation and causation – October 2015
Pairwise dependencies can be characterized using correlation but be aware that correlation only implies association, not causation. Conversely, causation implies association, not correlation.
Simple linear regression – November 2015
Linear regression is a flexible way to predict the values of one variable using the values of the other to find a ‘best line’ through the data points.

 We are looking for striking original images of the number 10 created by techniques or tools used for basic laboratory research in the biological sciences. This could be fluorescent cells patterned in the shape of a 10, a 10 written using two photon lithography, a DNA Origami-based 10, or 10 written using any number of other methods. The more imaginative the better!
We are looking for striking original images of the number 10 created by techniques or tools used for basic laboratory research in the biological sciences. This could be fluorescent cells patterned in the shape of a 10, a 10 written using two photon lithography, a DNA Origami-based 10, or 10 written using any number of other methods. The more imaginative the better!

