In 1995, Nature Genetics published a report by Eric Lander and Leonid Kruglyak, recommending clear statistical guidelines for reporting linkage results for complex traits. The paper had an immediate impact, setting the bar for what could or could not be called “significant” in the literature. Although originally focused on human genetic linkage studies, the guidelines set forth by Lander & Kruglyak influenced fields from model organism genetics to plant genetics, and eventually genome-wide association studies (GWAS).
In 1995, Nature Genetics published a report by Eric Lander and Leonid Kruglyak, recommending clear statistical guidelines for reporting linkage results for complex traits. The paper had an immediate impact, setting the bar for what could or could not be called “significant” in the literature. Although originally focused on human genetic linkage studies, the guidelines set forth by Lander & Kruglyak influenced fields from model organism genetics to plant genetics, and eventually genome-wide association studies (GWAS).
The mid-1990’s was a very exciting time in genetics. The human genome project had recently been announced and advances like microsatellite linkage maps of the human genome and multiplex sequencing technology were now available. Mapping genes underlying complex phenotypes was now a real possibility, and human geneticists were busy prospecting for genetic gold. However, as Lander & Kruglyak cautioned in their paper, the lack of clear guidelines could foster a spate a false positive reports that would, if left unchecked, discredit a the nascent field (for example, see this 1993 paper in Nature Genetics finding no evidence for a previously-reported linkage region for manic depressive illness).
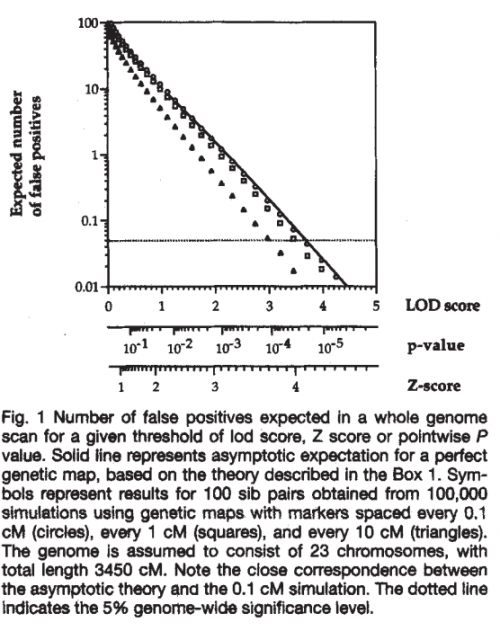
On the other hand, setting too high a bar for reporting significance would mean missing many true signals where they exist, an equally dangerous proposition for a new field. As explained in the paper, “striking the right balance requires both a mathematical understanding of how positive results will occur just by chance and a value judgment about the relative costs of false positives and false negatives.” The paper then outlines the mathematical and statistical arguments in favor of the standards we now all know and love.
I spoke with Leonid Kruglyak, co-author of this landmark paper, to get a sense of the context in which this paper came about, and the impact it had on the field at the time of publication. He first explained that it was finally possible to conduct genome-wide linkage studies with hundreds of individuals, allowing linkage mapping methods to be applied to complex traits (for example, this genome-wide screen for schizophrenia susceptibility genes published in the same issue). However, unlike Mendelian genes, there was no clue as to “how many signals there should be, or what their expected sizes were.” Thus, the need for a statistical framework.
This need was recognized as well by the Journal. As Prof Kruglyak recalls, Kevin Davies (founding editor of Nature Genetics) originally commissioned this work as a News & Views article, but it then evolved into a more extensive piece as its implications became clear. However, as he remembers, there was still a very strict deadline for the paper as it had to make the next issue (and these were still the days of hard-copy submissions). At the time, Prof Kruglyak was a young postdoc, so it fell to him to rush to the main FedEx office in downtown Boston before closing time, to make sure the manuscript got to the printer on time.
Prior to submitting the final text, Lander & Kruglyak produced some of the “original preprints”, sending a copy of the paper by snail mail or email to “everyone we knew in statistical genetics”, for comments and suggestions. After all, these guidelines would affect quite a lot of people and “signals that people would like to be results might not be real results anymore”.
Following publication, “the reactions came in essentially two flavors,” Prof Kruglyak recalls. There were those who thanked the authors, saying that someone really needed to do this. Others were less enthused. “They said, ‘you’re standing in the way of progress and making it harder to publish.’” In fact, Nature Genetics published two letters to the editor arguing that the proposed genome-wide significance threshold was too strict, or that at the very least additional discussion was warranted before these guidelines were adopted (see the letters here and here, and the authors’ reply here). Personally, I agree with the overall sentiment of Lander & Kruglyak as summed up in this portion of their reply: “The correspondents (all trained statisticians) argue that there is no need for guidelines because everyone should be able to interpret the genomewide significance of pointwise P values on their own. In our view, this is naïve. Most geneticists are not statisticians, and rules of thumb can be extremely helpful in promoting sensible discussion.”
The legacy of this paper is clear to anyone familiar with GWAS. “The GWAS community learned a lot from that whole experience [of false positive linkage reports],” says Prof Kruglyak. “There were many serious statistical geneticists involved [in the GWAS field] from the beginning, with a lot of carryover from the linkage era to the GWAS era.”
“Guidelines are not just ‘external gatekeepers’”, he noted. They are not just there to tell you what you can and can’t publish. “You know what they say, the easiest person to fool is yourself.” These guidelines were developed to help researchers understand their own findings better and decide which are worth following up. “You can often make up a plausible story, but how strong is the evidence?”